Abstract
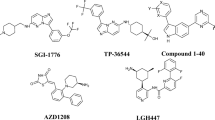
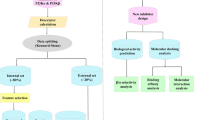
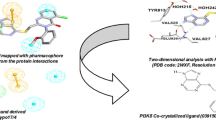
Pharmacophore modeling studies were undertaken for a series of compounds belonging several groups of phosphoinositide 3-kinase (PI3K) p110α inhibitors: 4-morpholino-2-phenylquinazolines derivatives, pyrido[3′,2′:4,5]furo-[3,2-d]pyrimidine derivatives, imidazo[1,2-a]pyridine derivatives, sulfonylhydrazone substituted imidazo[1,2-a]pyridines, and LY294002. A five-point pharmacophore with three hydrogen bond acceptors (A), one hydrophobic group (H), and one aromatic ring (R) as pharmacophore features was developed. The pharmacophore hypothesis yielded a statistically significant 3D-QSAR model, with a correlation coefficient of R 2 = 0.95 for training set compounds. The model generated showed excellent predictive power, with a correlation coefficient of Q 2 = 0.88 and r 2pret = 0.95 for a test set of 14 compounds. Furthermore, the structure–activity relationships of PI3K p110α inhibitors were elucidated and the activity differences between them discussed. Docking studies were also carried out wherein active and inactive compounds were docked into the active site of the PI3K p110α crystal structure to analyze PI3K p110α–inhibitor interactions. The results provide insights that will aid optimization of these classes of PI3K p110α inhibitors for better activity, and may prove helpful for further lead optimization and virtual screening.





Similar content being viewed by others
References
Fruman DA, Meyers RE, Cantley LC (1998) Annu Rev Biochem 67:481–507
Katso R, Okkenhaug K, Ahmadi K, White S, Timms J, Waterfield MD (2001) Annu Rev Cell Dev Biol 17:615–675
Djordjevic S, Driscoll PC (2002) Trends Biochem Sci 27:426–432
Jiang BH, Liu LZ (2008) BBA Proteins Proteomics 1784:150–158
Samuels Y, Wang Z, Bardelli A, Silliman N, Ptak J, Szabo S, Yan H, Gazdar A, Powell SM, Riggins GJ (2004) Science 304:554–571
Cantley LC, Neel BG (1999) Proc Natl Acad Sci USA 96:4240–4245
Graupera M, Guillermet-Guibert J, Foukas LC, Phng LK, Cain RJ, Salpekar A, Pearce W, Meek S, Millan J, Cutillas PR, Smith AJH, Ridley AJ, Ruhrberg C, Gerhardt H, Vanhaesebroeck B (2008) Nature 453:662–666
Hayakawa M, Kaizawa H, Moritomo H, Koizumi T, Ohishi T, Okada M, Ohta M, Tsukamoto S, Parker P, Workman P, Waterfield M (2006) Bioorg Med Chem 14:6847–6858
Hayakawa M, Kaizawa H, Moritomo H, Koizumi T, Ohishi T, Yamano M, Okada M, Ohta M, Tsukamoto S, Raynaud FI, Workman P, Waterfield MD, Parker P (2007) Bioorg Med Chem Lett 17:2438–2442
Hayakawa M, Kaizawa H, Kawaguchi K, Ishikawa N, Koizumi T, Ohishi T, Yamano M, Okada M, Ohta M, Tsukamoto S, Raynaud FI, Waterfield MD, Parker P, Workman P (2007) Bioorg Med Chem 15:403–412
Hayakawa M, Kawaguchi K, Kaizawa H, Koizumi T, Ohishi T, Yamano M, Okada M, Ohta M, Tsukamoto S, Raynaud FI, Parker P, Workman P, Waterfield MD (2007) Bioorg Med Chem 15:5837–5844
Frederick R, Denny WA (2008) J Chem Inf Model 48:629–639
Huang CH, Mandelker D, Schmidt-Kittler O, Samuels Y, Velculescu VE, Kinzler KW, Vogelstein B, Gabelli SB, Amzel LM (2007) Science 318:1744–1748
Phase 1.0 (2005) User manual. Schrodinger, New York
Dixon SL, Smondyrev AM, Knoll EH, Rao SN, Shaw DE, Friesner RA (2006) J Comput Aided Mol Des 20:647–671
Dixon SL, Smondyrev AM, Rao SN (2006) Chem Biol Drug Des 67:370–372
Evans DA, Doman TN, Thorner DA, Bodkin MJ (2007) J Chem Inf Model 47:1248–1257
Narkhede SS, Degani MS (2007) QSAR Comb Sci 26:744–753
Tawari NR, Bag S, Degani MS (2008) J Mol Model 14:911–921
Halgren TA, Murphy RB, Friesner RA, Beard HS, Frye LL, Pollard WT, Banks JL (2004) J Med Chem 47:1750–1759
Friesner RA, Banks JL, Murphy RB, Halgren TA, Klicic JJ, Mainz DT, Repasky MP, Knoll EH, Shelley M, Perry JK, Shaw DE, Francis P, Shenkin PS (2004) J Med Chem 47:1739–1749
Sherman W, Day T, Jacobson MP, Friesner RA, Farid R (2006) J Med Chem 49:534–553
Halgren TA (1996) J Comput Chem 17:520
MacroModel 2.0 (2006) User manual. Schrodinger, New York
Tropsha A (2005) In: Oprea TI (ed) Chemoinformatics in drug discovery. Wiley, Weinheim, pp 437–455
Knight ZA, Gonzalez B, Feldman ME, Zunder ER, Goldenberg DD, Williams O, Loewith R, Stokoe D, Balla A, Toth B, Balla T, Weiss WA, Williams RL, Shokat KM (2006) Cell 125:733–747
Zvelebil MJ, Waterfield MD, Shuttleworth SJ (2008) Arch Biochem Biophys 477:404–410
Amzel LM, Huang CH, Mandelker D, Lengauer C, Gabelli SB, Vogelstein B (2008) Nat Rev Cancer 8:665–669
Acknowledgment
This work was financially supported by Natural Science Foundation of Shaanxi Province (NO. SJ08C207).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, Y., Wang, Y. & Zhang, F. Pharmacophore modeling and 3D-QSAR analysis of phosphoinositide 3-kinase p110α inhibitors. J Mol Model 16, 1449–1460 (2010). https://doi.org/10.1007/s00894-010-0659-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-010-0659-y