Summary
Background
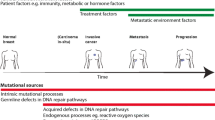
A comprehensive and consistent picture of the genetic changes that underlie breast cancer initiation, development, and progression remains unresolved. The MCF10 series of cell lines represents many steps in that progression. We performed high resolution mapping of the MCF10 series of cell lines to identify specific gene targets to elucidate the molecular correlates of immortalization, development, and progression of breast cancer at the level of individual genes.
Design
We evaluated the initial untransformed outgrowths (MCF-10MS and MCF-10A) with six transformed cell lines with benign proliferations (MCF-10AT1, MCF-10AT1kcl2), carcinoma in situ (MCF-10CA1h cl13), and invasive carcinoma (MCF-10CA1h cl2, MCF-10CA1a cl1, MCF-10CA1d cl1). Losses and gains of loci at 112 unique human genome sites were interrogated using the multiplex ligation-dependent probe amplification assay (MLPA).
Results
Cytogenetic alterations in the four benign progenitors that persisted in the CIS and invasive cell lines corresponded to gains and losses of genes by MLPA. MCF-10MS had only normal gene copies. The untransformed MCF-10A had cytogenetic gain of 5q13-qter with corresponding gains of the IL3, IL4 and IL12B genes at 5q31-q33; gain of distal 19q12-qter was reflected in gains in KLK3 and BAX gene loci at 19q13-q13.4. The observed genic gain of cMYC at 8q24.12 was not indicated by cytogenetics. The apparently balanced t(3;9) component of the t(3;9)(p13;p22)t(3;5)(p26;q31) resulted in complete loss of the CDKN2A and CDKN2B genes at 9p21. Additional clonal cytogenetic changes in the DCIS cell line (MCF-10A1h cl13) involving chromosomes 1, 3 and 10 persisted in the invasive progeny, with gain of corresponding genes at 1p13 (BCAR2, BCAR3, NRAS, TGFB2), at 3p12–13 (IL12A), and 3q21–27 (MME, PIK3CA, BCL6).
Conclusions
Our study adopted a comprehensive exploration of genetic changes using high resolution molecular probes applied to the MCF10 family of cell lines to identify individual genes in a continuum starting from normal breast epithelial cells and progressing through immortalization, transformation and invasive malignancy. Homozygous loss of CDKN2A and CDKN2B genes and gain of MYC were initiating immortalization events. Transformation and progression to malignancy event were marked by gains of IL13, VEGF, HRAS, TRAF2, and BCAS2, IL12A, and MME, respectively.
Similar content being viewed by others
References
Soule HD, Maloney TM, Wolman SR, Peterson WD, Jr., Brenz R, McGrath CM, Russo J, Pauley RJ, Jones RF, Brooks SC, Isolation and characterization of a spontaneously immortalized human breast epithelial cell line, MCF-10 Cancer Res 50:6075–6086, 1990
Wolman SR, Mohamed AN, Heppner GH, Soule HD, Chromosomal markers of immortalization in human breast epithelium Genes Chromosomes Cancer 10:59–65, 1994
Basolo F, Elliott J, Tait L, Chen XQ, Maloney T, Russo IH, Pauley R, Momiki S, Caamano J, Klein-Szanto AJ, et al. Transformation of human breast epithelial cells by c-Ha-ras oncogene Mol Carcinog 4:25–35, 1991
Miller FR, Soule HD, Tait L, Pauley RJ, Wolman SR, Dawson PJ, Heppner GH, Xenograft model of progressive human proliferative breast disease J Natl Cancer Inst 85:1725–1732, 1993
Dawson PJ, Wolman SR, Tait L, Heppner GH, Miller FR, MCF10AT: a model for the evolution of cancer from proliferative breast disease Am J Pathol 148:313–319, 1996
Iravani S, Mora L, Miller FR, Dawson PJ, Altered expression of c-erbB-2, DF3, b72.3, p53 and Ki-67 with progression and differentiation to two distinct histologic types of invasive carcinoma in the MCF10AT human xenograft model of proliferative breast disease Int J Oncol 12:369–375, 1998
Shekhar MP, Nangia-Makker P, Wolman SR, Tait L, Heppner GH, Visscher DW, Direct action of estrogen on sequence of progression of human preneoplastic breast disease Am J Pathol 152:1129–1132, 1998
Strickland LB, Dawson PJ, Santner SJ, Miller FR, Progression of premalignant MCF10AT generates heterogeneous malignant variants with characteristic histologic types and immunohistochemical markers Breast Cancer Res Treat 64:235–240, 2000
Miller FR, Santner SJ, Tait L, Dawson PJ, MCF10DCIS.com xenograft model of human comedo ductal carcinoma in situ J Natl Cancer Inst 92:1185–1186, 2000
Santner SJ, Dawson PJ, Tait L, Soule HD, Eliason J, Mohamed AN, Wolman SR, Heppner GH, Miller FR, Malignant MCF10CA1 cell lines derived from premalignant human breast epithelial MCF10AT cells Breast Cancer Res Treat 65:101–110, 2001
Worsham MJ, Pals G, Schouten JP, Van Spaendonk RM, Concus A, Carey TE, Benninger MS, Delineating genetic pathways of disease progression in head and neck squamous cell carcinoma Arch Otolaryngol Head Neck Surg 129:702–708, 2003
Schouten JP, McElgunn CJ, Waaijer R, Zwijnenburg D, Diepvens F, Pals G, Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification Nucleic Acids Res 30:e57, 2002
Lisitsyn N, Wigler M, Cloning the differences between two complex genomes Science 259:946–951, 1993
Rouse J, Jackson SP, Interfaces between the detection, signaling, and repair of DNA damage Science 297:547–551, 2002
Kolodner RD, Putnam CD, Myung K, Maintenance of genome stability in Saccharomyces cerevisiae Science 297:552–557, 2002
Maser RS, DePinho RA, Connecting chromosomes, crisis, and cancer Science 297:565–569, 2002
Caspersson T, Farber S, Foley GE, Kudynowski J, Modest EJ, Simonsson E, Wagh U, Zech L, Chemical differentiation along metaphase chromosomes Exp Cell Res 49:219–222, 1968
Trask BJ, 1999 Genome Analysis: A Laboratory Manual New York, Cold Spring Harbor Laboratory Press
Mitelman F, 1998 Catalog of Chromosome Aberrations in Cancer New York, Wiley
Collins FS, Positional cloning moves from perditional to traditionalNat Genet 9:347–350, 1995
Pinkel D, Landegent J, Collins C, Fuscoe J, Segraves R, Lucas J, Gray J, Fluorescence in situ hybridization with human chromosome-specific libraries: detection of trisomy 21 and translocations of chromosome 4 Proc Natl Acad Sci USA85:9138–9142, 1988
Albertson DG, Ylstra B, Segraves R, Collins C, Dairkee SH, Kowbel D, Kuo WL, Gray JW, Pinkel D, Quantitative mapping of amplicon structure by array CGH identifies CYP24 as a candidate oncogene Nat Genet 25:144–146, 2000
Hayashizaki Y, Hirotsune S, Okazaki Y, Hatada I, Shibata H, Kawai J, Hirose K, Watanabe S, Fushiki S, Wada S, et al. Restriction landmark genomic scanning method and its various applications Electrophoresis 14:251–258, 1993
Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, Waldman F, Pinkel D, Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors Science 258:818–821, 1992
Ginzinger DG, Godfrey TE, Nigro J, Moore DH, 2nd, Suzuki S, Pallavicini MG, Gray JW, Jensen RH, Measurement of DNA copy number at microsatellite loci using quantitative PCR analysis Cancer Res 60:5405–5409, 2000
Hodgson G, Hager JH, Volik S, Hariono S, Wernick M, Moore D, Nowak N, Albertson DG, Pinkel D, Collins C, Hanahan D, Gray JW, Genome scanning with array CGH delineates regional alterations in mouse islet carcinomas Nat Genet 29:459–464, 2001
Snijders AM, Nowak N, Segraves R, Blackwood S, Brown N, Conroy J, Hamilton G, Hindle AK, Huey B, Kimura K, Law S, Myambo K, Palmer J, Ylstra B, Yue JP, Gray JW, Jain AN, Pinkel D, Albertson DG, Assembly of microarrays for genome-wide measurement of DNA copy number Nat Genet 29:263–264, 2001
Pollack JR, Sorlie T, Perou CM, Rees CA, Jeffrey SS, Lonning PE, Tibshirani R, Botstein D, Borresen-Dale AL, Brown PO, Microarray analysis reveals a major direct role of DNA copy number alteration in the transcriptional program of human breast tumors Proc Natl Acad Sci USA 99:12963–12968, 2002
Hyman E, Kauraniemi P, Hautaniemi S, Wolf M, Mousses S, Rozenblum E, Ringner M, Sauter G, Monni O, Elkahloun A, Kallioniemi OP, Kallioniemi A, Impact of DNA amplification on gene expression patterns in breast cancer Cancer Res 62:6240–6245, 2002
Schrock E, du Manoir S, Veldman T, Schoell B, Wienberg J, Ferguson-Smith MA, Ning Y, Ledbetter DH, Bar-Am I, Soenksen D, Garini Y, Ried T, Multicolor spectral karyotyping of human chromosomes Science 273:494–497, 1996
Haber DA, Splicing into senescence: the curious case of p16 and p19ARF Cell 91:555–558, 1997
Johnson DG, Walker CL, Cyclins and cell cycle checkpoints Annu Rev Pharmacol Toxicol 39:295–312, 1999
Esteve A, Martel-Planche G, Sylla BS, Hollstein M, Hainaut P, Montesano R, Low frequency of p16/CDKN2 gene mutations in esophageal carcinomas Int J Cancer 66:301–304, 1996
Xing EP, Nie Y, Song Y, Yang GY, Cai YC, Wang LD, Yang CS, Mechanisms of inactivation of p14ARF, p15INK4b, and p16INK4a genes in human esophageal squamous cell carcinoma Clin Cancer Res 5:2704–2713, 1999
Gamieldien W, Victor TC, Mugwanya D, Stepien A, Gelderblom WC, Marasas WF, Geiger DH, van Helden PD, p53 and p16/CDKN2 gene mutations in esophageal tumors from a high-incidence area in South Africa Int J Cancer 78:544–549, 1998
Hannon GJ, Beach D, p15INK4B is a potential effector of TGF-beta-induced cell cycle arrest Nature 371:257–261, 1994
Gronbaek K, de Nully Brown P, Moller MB, Nedergaard T, Ralfkiaer E, Moller P, Zeuthen J, Guldberg P, Concurrent disruption of p16INK4a and the ARF-p53 pathway predicts poor prognosis in aggressive non-Hodgkin’s lymphoma Leukemia 14:1727–1735, 2000
Bardeesy N, Morgan J, Sinha M, Signoretti S, Srivastava S, Loda M, Merlino G, DePinho RA, Obligate roles for p16(Ink4a) and p19(Arf)-p53 in the suppression of murine pancreatic neoplasia Mol Cell Biol 22:635–643, 2002
Lukas J, Aagaard L, Strauss M, Bartek J, Oncogenic aberrations of p16INK4/CDKN2 and cyclin D1 cooperate to deregulate G1 control Cancer Res 55:4818–4823, 1995
Yokota T, Yoshimoto M, Akiyama F, Sakamoto G, Kasumi F, Nakamura Y, Emi M, Frequent multiplication of chromosomal region 8q24.1 associated with aggressive histologic types of breast cancers Cancer Lett 139:7–13, 1999
Tsuneizumi M, Emi M, Nagai H, Harada H, Sakamoto G, Kasumi F, Inoue S, Kazui T, Nakamura Y, Overrepresentation of the EBAG9 gene at 8q23 associated with early-stage breast cancers Clin Cancer Res 7:3526–3532, 2001
Sieburth D, Jabs EW, Warrington JA, Li X, Lasota J, LaForgia S, Kelleher K, Huebner K, Wasmuth JJ, Wolf SF, Assignment of genes encoding a unique cytokine (IL12) composed of two unrelated subunits to chromosomes 3 and 5 Genomics 14:59–62, 1992
Sattler HP, Lensch R, Rohde V, Zimmer E, Meese E, Bonkhoff H, Retz M, Zwergel T, Bex A, Stoeckle M, Wullich B, Novel amplification unit at chromosome 3q25–q27 in human prostate cancer Prostate 45:207–215, 2000
Liu YJ, Johnson GD, Gordon J, MacLennan IC, Germinal centres in T-cell-dependent antibody responses Immunol Today 13:17–21, 1992
Rowe M, Rowe DT, Gregory CD, Young LS, Farrell PJ, Rupani H, Rickinson AB, Differences in B cell growth phenotype reflect novel patterns of Epstein-Barr virus latent gene expression in Burkitt’s lymphoma cells Embo J 6:2743–2751, 1987
Cutrona G, Leanza N, Ulivi M, Melioli G, Burgio VL, Mazzarello G, Gabutti G, Roncella S, Ferrarini M, Expression of CD10 by human T cells that undergo apoptosis both in vitro and in vivo Blood 94:3067–3076, 1999
Nagasaki K, Maass N, Manabe T, Hanzawa H, Tsukada T, Kikuchi K, Yamaguchi K, Identification of a novel gene, DAM1, amplified at chromosome 1p13.3–21 region in human breast cancer cell lines Cancer Lett 140:219–226, 1999
Maass N, Rosel F, Schem C, Hitomi J, Jonat W, Nagasaki K, Amplification of the BCAS2 gene at chromosome 1p13.3–21 in human primary breast cancer Cancer Lett 185:219–223, 2002
Forozan F, Veldman R, Ammerman CA, Parsa NZ, Kallioniemi A, Kallioniemi OP, Ethier SP, Molecular cytogenetic analysis of 11 new breast cancer cell lines Br J Cancer 81:1328–1334, 1999
Zimonjic DB, Keck-Waggoner CL, Yuan BZ, Kraus MH, Popescu NC, Profile of genetic alterations and tumorigenicity of human breast cancer cells Int J Oncol 16:221–230, 2000
Forozan F, Mahlamaki EH, Monni O, Chen Y, Veldman R, Jiang Y, Gooden GC, Ethier SP, Kallioniemi A, Kallioniemi OP, Comparative genomic hybridization analysis of 38 breast cancer cell lines: a basis for interpreting complementary DNA microarray data Cancer Res 60:4519–4525, 2000
Fearon ER, Vogelstein B, A genetic model for colorectal tumorigenesis Cell 61:759–767, 1990
Nowell PC, The clonal evolution of tumor cell populations Science 194:23–28, 1976
Frame S, Balmain A, Integration of positive and negative growth signals during ras pathway activation in vivo Curr Opin Genet Dev 10:106–113, 2000
Marx J, Debate surges over the origins of genomic defects in cancerScience 297:544–546, 2002
Brown K, Strathdee D, Bryson S, Lambie W, Balmain A, The malignant capacity of skin tumours induced by expression of a mutant H-ras transgene depends on the cell type targeted Curr Biol 8:516–524, 1998
Liotta L, Petricoin E, Molecular profiling of human cancer Nat Rev Genet 1:48–56, 2000
Acknowledgements
Supported by in part by DOD Grants, DAMD DAMD17-00-1-0288 (MJW), DAMD17-02-1-0406 (MJW), and NIH Grants CA70923 (MJW), and DE15990 (MJW)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Worsham, M.J., Pals, G., Schouten, J.P. et al. High-resolution mapping of molecular events associated with immortalization, transformation, and progression to breast cancer in the MCF10 model. Breast Cancer Res Treat 96, 177–186 (2006). https://doi.org/10.1007/s10549-005-9077-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-005-9077-8