Abstract
The heat-shock response is an evolutionarily conserved cellular defense mechanism against environmental stresses, characterized by the rapid synthesis of heat-shock proteins (HSPs). HSP70, a highly inducible molecular chaperone, assists in refolding or clearance of damaged proteins, thereby having a central role in maintaining intracellular homeostasis and thermotolerance. To date, induction of HSP70 expression has been described extensively at the transcriptional level. However, post-translational regulation of HSP70, such as protein stability, is only partially understood. In this study, we investigated the role of OLA1 (Obg-like ATPase 1), a previously uncharacterized cytosolic ATPase, in regulating the turnover of HSP70. Downregulation of OLA1 in mammalian cells by either RNAi or targeted gene disruption results in reduced steady-state levels of HSP70, impaired HSP70 induction by heat, and functionally, increased cellular sensitivity to heat shock. Conversely, overexpression of OLA1 correlates with elevated HSP70 protein levels and improved thermal resistance. Protein–protein interaction assays demonstrated that binding of OLA1 to the HSP70 carboxyl terminus variable domain hinders the recruitment of CHIP (C-terminus of Hsp70-binding protein), an E3 ubiquitin ligase for HSP70, and thus prevents HSP70 from the CHIP-mediated ubiquitination. These findings suggest a novel molecular mechanism by which OLA1 stabilizes HSP70, leading to upregulation of HSP70 as well as increased survival during heat shock.
Similar content being viewed by others
Main
The-heat-shock response is an evolutionarily conserved cellular defense mechanism against environmental challenges, such as heat shock, oxidative stress, heavy metals, and toxins.1, 2, 3, 4 Following stress exposure, a family of heat-shock transcription factors (HSFs) rapidly induces the expression of cytoprotective heat-shock proteins (HSPs) including the HSP70 molecular chaperone.5 HSP70 catalyzes the proper folding of nascent proteins, and under stressful conditions, refolds the damaged proteins, or targets unrepairable proteins for degradation.6 HSP70 has critical roles in protein homeostasis and thermotolerance in mammalian cells,7, 8, 9 and also participates in multiple cellular signaling events and developmental processes.8, 10 HSP70 has been implicated in several neurodegenerative diseases related to protein misfolding and aggregation, such as Alzheimer’s disease, Parkinson’s disease, and Huntington’s disease.11, 12, 13, 14 Additionally, HSP70 is also found highly expressed in various malignant tumors with its expression positively correlated with tumorigenesis, metastasis, and poor therapeutic outcome in human cancers.15, 16, 17
Owing to its important basal and stress-response functions, HSP70 expression is tightly controlled within a cell, and both HSP70 loss and gain have been shown to contribute to multiple dysfunctional or pathological processes.11, 18 Upregulation of HSP70 expression is largely attributed to the binding of HSF1 to the heat-shock element (HSE) located at the promoter region of the heat-shock gene (HSP70A1A).19 However, various post-transcriptional, translational, and post-translational mechanisms also contribute to the regulation of HSP70 expression in both steady-state and heat-shock response conditions.20, 21 C-terminus of Hsp70-binding protein (CHIP), an E3 ligase, is shown to mediate ubiquitin-dependent degradation of HSP70-bound protein substrates as well as HSP70 itself,18, 22 representing an important mechanism that shuts down the HSP70 response after the damaged proteins are removed. This finding particularly emphasizes the importance of protein degradation in determining the overall levels of the HSP before, during, and after the heat shock.
In our recent efforts in search of novel cellular factors that modulate protein turnover, HSP70 was found to interact with OLA1 (Obg-like ATPase 1), a newly identified ATPase member of the YchF subfamily GTPases. GTPases are the most abundant class of NTP-binding proteins,23 and have a crucial role in the regulation of diverse cellular processes such as protein translation, intracellular transport, signal transduction, and growth.24, 25 YchF proteins are highly conserved from yeast to human,23, 26 and unlike other GTPases, bind and hydrolyze ATP more efficiently than GTP.27 However, physiological functions of YchF proteins, especially human OLA1, are poorly understood. The yeast YBR025c/Ola1 is associated with the 26S proteasome,28, 29 whereas the protozoan TcYchF/Ola1 interacts with the proteasome subunit RPN10, ribosomal subunits, and polysomes,30 suggesting a function in the regulation of protein translation and degradation. Sun et al.31 reported that OLA1 is downregulated under DNA damage stress, such as etoposide, adriamycin, UV, and ionizing radiation. Recently, our group demonstrated that OLA1 functions as a negative regulator in the antioxidant response by transcription-independent mechanisms.32, 33 Therefore, it is plausible to reason that OLA1 may function in the regulation of multiple stress responses and, particularly, the classical physical stresses such as heat shock. In the present study, we demonstrate that OLA1 protects cells from heat shock by interfering with the binding and function of the E3 ligase CHIP to HSP70, leading to stabilization of HSP70. Our findings pinpoint to a novel post-translational mechanism by which HSP70 is upregulated due to reduced degradation and increased stability.
Results
OLA1 protects cells from heat shock-induced cell death
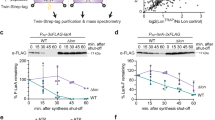
To assess the potential involvement of OLA1 in control of the heat-shock response, Ola1-knockout (−/−) and wild-type (+/+) mouse embryonic fibroblast cells (MEFs) were treated with heat shock at either 42 or 45°C, and their survival examined following a 6-h recovery period. Compared with Ola1+/+ MEFs, Ola1−/− cells were more sensitive to heat shock at both temperatures as evidenced by morphological changes (Figure 1a) and significantly decreased cell viability (Figure 1b). To examine if this phenotype is cell type or species dependent, we generated shRNA-mediated OLA1 knockdown cells derived from human breast cancer cell line MDA-MB-231. Cells stably transfected with OLA1 shRNA exhibited diminished OLA1 expression and were significantly more sensitive to heat shock compared with control cells transfected with non-targeting shRNA (Supplementary Figures S1a–c). Further, we tested whether overexpression of OLA1 could protect cells from heat shock. Human embryonic kidney (HEK) 293T cells transiently transfected with a YFP-OLA1 fusion construct exhibited significantly increased survival in response to heat shock (Figures 1d and e) compared with control cells expressing the YFP construct. These results suggest that OLA1 may function to protect cells from heat shock-induced cell death.
OLA1 protects mammalian cells from heat shock-induced cell death. (a) Effects of heat shock on mouse embryonic fibroblasts (MEFs). Primary cultures of wild-type (WT) and Ola1−/− (OLA1 KO) MEFs prepared from 12.5-day embryos (⩽ passage 5) were grown at 37°C (NT) or incubated at either 42°C (heat shock, HS) for 1 h or 45°C (lethal dose heat shock, LHS) for 40 min, and allowed to recover for 6 h at 37°C. Morphological changes were observed with a microscope. Representative images from three independent experiments are shown. (b) Ola1−/− MEFs exhibited decreased cell viability in response to heat shock. Primary MEFs were either untreated (NT) or treated with heat (42°C, HS) for 1 h and recovered at 37°C for 6 h. Cell viability was measured by MTS assay (n=3 independent experiments). (c) Deficiency of Ola1 protein level in Ola1-knockout MEFs was confirmed by western blot analysis. β-Actin served as a loading control. (d) Overexpression of OLA1 protects cells from heat shock. HEK-293T cells were transfected with the YFP-OLA1 or YFP constructs as described in Materials and Methods. Forty-eight hours after transfection, the cells were exposed to either HS for 1 h or an LHS for 40 min. Following a recovery of 6 h at 37°C, cellular morphologies were observed (n=2 independent transfections). (e) Quantitative analysis of cell viability loss due to heat shock. HEK-293T cells were transfected with YFP-OLA1 or YFP constructs, and treated as in (b). MTS assay was used to determine the cell viability. Mean viability is shown. Error bars represent the standard deviations of at least triplicate samples. **P<0.01. (Significance was determined by the two-tailed Student’s t-test.)
Since the above observations were made after a period of heat-shock recovery, reflecting both acute stress response and chronic recovery, we sought to further examine the dynamic course of an acute phase of heat-shock response in OLA1-deficient and overexpressing cells. In contrast to Ola1+/+ MEFs, which were still viable after a 3-h acute heat shock, cell death was observed in Ola1−/− MEFs as early as 60 min following the heat shock (Figures 2a and c). Similarly, cell death induction by heat shock during the acute phase was significantly higher in MDA-MB-231 cells with stable knockdown of OLA1 compared with control cells (Figures 2b and d). Conversely, overexpression of OLA1 in HEK-293T cells decreased cell death induction from acute heat shock (Supplementary Figures S2a and b). Taken together, our findings suggest that OLA1 protects cells from heat shock in both the acute and chronic phases of stress response, and that this cell survival-promoting function of OLA1 is not limited to a specific type of cells.
OLA1 protects cells from acute phase heat shock. (a) Dynamics of the acute phase of heat-shock response in Ola1-knockout MEFs. Immortalized wide-type (WT) and Ola1−/− MEFs, generated by repetitive subculturing of the primary MEFs for >30 passages, were grown at 37°C (NT) or heat shocked at 42°C for various durations as indicated. Representative microscopic images from three independent experiments are shown. (b) Dynamics of the acute phase of heat-shock response in stable OLA1-knockdown MDA-MB-231 cells. Cells were incubated at 37°C (NT) or exposed to heat shock at 42°C for various durations as indicated, and morphological changes were observed under a microscope (n=2 independent experiments). (c) Quantitative analysis of cell viability in the acute phase of heat-shock response. MTS assays were applied to MEFs receiving heat-shock treatment as described in (a). (d) MTS analysis of the OLA1-knockdown MDA-MB-231 cells under heat shock as described in (b). Error bars represent the standard deviations of at least triplicate samples. **P<0.01. (Significance was determined by the two-tailed Student’s t-test.)
OLA1 deficiency affects basal and heat shock-inducible HSP70
We investigated if the observed loss of thermal resistance in OLA1-deficient cells was due to insufficient HSPs or impaired induction of their expression after heat shock. We sought to assess the levels of HSP70, a major HSP, in Ola1-knockout MEFs. HSP70 protein levels were significantly lower in Ola1−/− MEFs compared with Ola1+/+ MEFs (Figure 3a). To further confirm these findings, we examined HSP70 levels in human cell lines with RNA interference-mediated knockdown of OLA1 expression. Stable OLA1-knockdown MDA-MB-231 cells exhibited decreased basal HSP70 levels compared with their control counterparts (Figure 3b). Moreover, the role of OLA1 in HSP70 expression was verified in transient transfection experiments. Transient overexpression of OLA1 in either HEK-293T or MDA-MB-231 cells resulted in increased HSP70 protein expression, whereas transient knockdown of OLA1 in these cells led to decreased HSP70 levels (Supplementary Figures S3a–d). It is worthwhile to note that basal levels of other HSPs, including HSP110, HSP90, and HSP27, were not affected by the knockdown of OLA1 in either HEK-293T or MDA-MB-231 cells (Supplementary Figures S3e and f). To test the inducibility of HSP70 in OLA1-deficient cells, a non-lethal heat shock was applied and the cells were allowed to recover for 6 h. HSP70 protein levels were significantly lower in Ola1−/− MEFs relative to Ola1+/+ MEFs (Figure 3c), as well as in MDA-MB-231 cells with stable OLA1 knockdown relative to their control counterparts (Figure 3d). These results demonstrate that OLA1 positively regulates both the steady-state and heat shock-inducible levels of HSP70.
Basal and inducible levels of HSP70 protein in OLA1-deficient cells. (a) HSP70 protein expression in immortalized Ola1+/+ and Ola1−/− MEFs was evaluated by western blot analysis using anti-HSP70 antibody. Anti-OLA1 antibody was also used to confirm the absence of Ola1 in the Ola1−/− MEFs. A non-specific band (NS) serves as the loading control. Relative HSP70 band intensities from the scanned western blot are shown in the right panel. Data are presented as mean±S.D. values from three independent experiments. **P<0.01 as determined by the two-tailed Student’s t-test. (b) Western blot analysis of HSP70 protein in stable OLA1-knockdown MDA-MB-231 cells. β-Actin was used as the loading control. Relative HSP70 band intensities from the scanned western blot are shown in the right panel. Data are presented as mean±S.D. values from three independent experiments. *P<0.05 as determined by the two-tailed Student’s t-test. (c) Heat induction of HSP70 protein is impaired in Ola1−/− MEFs. Ola1+/+ and Ola1−/− MEFs were grown at 37°C (NT) or exposed to heat shock (HS, 42°C) for 1 h. After 6 h recovery at 37°C, cell lysates were collected for western blot analysis. β-Actin was used as the loading control. The bar graph on the right represents the quantification of levels of HSP70 expression in the HS group relative to that in the NT group (fold induction). The bars are the means±S.D. values for three independent experiments. **P<0.01 as determined by the two-tailed Student’s t-test. (d) Reduced induction of HSP70 protein in stable OLA1-knockdown MDA-MB-231 cells after heat shock (42°C, 1 h). β-Actin was used as the loading control. The bar graph represents the quantification of levels of HSP70 expression in the HS group relative to that in the NT group (fold induction). The bars are the means±S.D. values for three independent experiments. **P<0.01 as determined by the two-tailed Student’s t-test
We next questioned whether OLA1 regulates HSP70 at the transcriptional level. Quantitative RT-PCR analysis revealed that there were no significant differences in HSP70 mRNA expression following transient knockdown of OLA1 in MDA-MB-231 and HEK-293T cells (Supplementary Figures S4a and b). Similarly, we did not find significant differences in HSP70 mRNA levels between Ola1+/+ and Ola1−/− MEFs or between MDA-MB-231 cells with stable knockdown of OLA1 and their control counterparts (Supplementary Figures S4c and d). In addition, knockout or knockdown of OLA1 did not have a significant effect on the induction of HSP70 mRNA following heat shock, as we noted similar increase rates in the mRNA levels among all isogenic cell line pairs tested (Supplementary Figures S4e and f). Furthermore, using a luciferase reporter system with an innate human HSP70 promoter, we found that OLA1 deficiency did not significantly alter the heat-induced luciferase activities (Supplementary Figure S5). These findings suggest that OLA1-mediated changes in the expression of HSP70 are unlikely caused by OLA1’s action at the transcriptional level.
OLA1 binds with HSP70
The discordant changes in the protein and transcript levels of HSP70 prompted us to further explore the mechanism by which OLA1 increases HSP70 protein expression, using protein–protein interaction assays to determine whether OLA1 binds to HSP70. Immunoprecipitation followed by western blot analysis in HEK-293T cells demonstrated that HSP70 interacts with ectopically expressed FLAG-tagged OLA1 (Figure 4a) as well as the endogenous OLA1 (Figure 4b). Similar results were obtained when reciprocal immunoprecipitation was performed (Figure 4c). This endogenous OLA1–HSP70 interaction was also observed in additional cell lines (HeLa and MDA-MB-231 cells; Figures 4d and e). Furthermore, in vitro binding assays using recombinant proteins revealed the direct binding of OLA1 and HSP70 (Figure 4f). On the other hand, immunofluorescence staining followed by confocal microscopy analysis demonstrated that OLA1 was co-localized with HSP70 in the cytoplasm in both HEK-293T and HeLa cells (Figures 4g and h). In an effort to screen for protein interaction partners of OLA1, we performed immunoprecipitation/mass spectrometry analysis of HEK-293T cells that were transfected with a Flag-OLA1-expressing construct and immunoprecipitated with anti-FLAG antibody, and found that HSP70 was among the OLA1-interacting proteins. These results strongly suggest that OLA1 directly interacts with HSP70 under physiological conditions.
OLA1 binds to HSP70 both in vivo and in vitro. (a) Ectopically expressed OLA1 binds with intracellular HSP70. HEK-293T cells were transfected with a control vector or expression vector encoding Flag-OLA1. Forty-eight hours after the transfection, the cell lysates were immunoprecipitated with anti-FLAG antibody and immunoblotted with both anti-Flag and anti-HSP70 antibodies (upper panel). The abundance of HSP70 and Flag-OLA1 in the whole cell lysates was also evaluated by the same antibodies (lower panel). (b and c) Binding of endogenously expressed OLA1 and HSP70 in HEK-293T cells. OLA1 protein in HEK-293T cell lysates was immunoprecipitated with anti-OLA1 antibody and the precipitates were analyzed for HSP70 by immunoblotting (b). Reciprocally, cell lysates from HEK-293T cells were immunoprecipitated with anti-HSP70 antibody and immunoblotted with both anti-HSP70 and anti-OLA1 (c). In both cases, the whole cell lysates were also analyzed by anti-HSP70 and OLA1 antibodies. (d and e) OLA1 binds to HSP70 endogenously in both HeLa and MDA-MB-231 cells. Endogenous OLA1 was immunoprecipitated in HeLa (d) and MDA-MB-231 (e) cells, respectively. OLA1 protein in the cell lysates was immunoprecipitated with anti-OLA1 antibody and the precipitates were analyzed by immunoblotting with anti-HSP70 antibody. (f) OLA1 binds with HSP70 in vitro. Recombinant human HSP70 and OLA1 proteins were co-incubated at 4°C in binding buffer for 3 h, and immunoprecipitated with anti-HSP70 antibody. The precipitates were then analyzed by immunoblotting with anti-OLA1 antibody (upper panel). The ‘input’ HSP70 and OLA1 proteins were verified by Coomassie blue staining (lower panel). (g and h) Intracellular co-localization of OLA1 and HSP70. HEK-293T (g) or HeLa (h) cells were grown and fixed on sterile glass coverslips, stained with anti-OLA1 (green) and anti-HSP70 (red) antibodies, and counterstained with Hoechst 33342 (blue) for the nuclei. Single color images were taken with a Confocal microscope, and superimposed by the FC-10-ASW 3.0 software. Scale bar: 10 μm
OLA1 stabilizes HSP70
Based on the above results, we hypothesized that the direct binding of OLA1 to HSP70 may have a role in the stability of HSP70 protein. The turnover rate of HSP70 in Ola1−/− and the wild-type MEFs was examined by incubating cells in medium containing the protein synthesis inhibitor cycloheximide. Interestingly, we found that the half-life time of HSP70 protein in Ola1-deficient MEFs (∼30 min) is shorter than that in the wild-type MEFs (>60 min) (Figure 5a). A similarly shortened HSP70 half-life time was also observed in MDA-MB-231 cells with stable knockdown of OLA1 (Figure 5b). These results indicate that OLA1 binding stabilizes HSP70 protein. Moreover, when a Myc-tagged HSP70 overexpressing construct was introduced into both the control MDA-MB-231 cells and cells with stable knockdown of OLA1, a lower level of Myc-HSP70 protein was detected in the latter cell line (Figure 5c). However, when an unrelated construct (containing HA-USP4) was co-transfected, the difference in expression levels of this reference protein in these two types of cells was insignificant (Figure 5c). These results further support the hypothesis that OLA1 stabilizes HSP70.
OLA1 stabilizes HSP70. (a and b) The half-life of HSP70 protein is shortened in OLA1-deficient cells. MEFs (a) and MDA-MB-231 cells stably transfected with OLA1-shRNA (b) were treated with cycloheximide (CHX) for 0, 30, 60, and 120 min, and probed for HSP70 or OLA1 on western blots. β-Actin was used as the loading control. Quantitative analyses of the relative HSP70 protein levels are shown as curve graphs in the right panels. The levels of HSP70 at 0 min of CHX treatment are designated as ‘1’. Data are presented as mean±S.D. values from three independent experiments. The mean values were fitted to exponential decay curves to estimate half-life (t1/2). *P<0.05, **P<0.01. (Significance was determined by the two-tailed Student’s t-test.) (c) In vitro transfection assay. OLA1 stable knockdown MDA-MB-231 cells and control cells were co-transfected with the Myc-HSP70 and HA-USP4 constructs, or with the corresponding empty vectors. Forty-eight hours after the transfection, cell lysates were immunoblotted with anti-Myc, anti-HA, anti-OLA1, and anti-β-actin antibodies. β-Actin served as a loading control (n=2 independent transfections)
OLA1 binds with HSP70 C-terminal variable domain
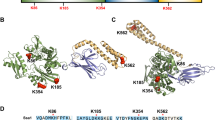
It was recently reported that HSP70 stability is controlled by the ubiquitin-proteasome system (UPS).22 CHIP, an E3 ligase of HSP70, has been shown to ubiquitinate HSP70 and promote HSP70 degradation under both steady-state and heat-shock response conditions.18, 22 Since we found that OLA1 modulates levels of basal and heat-induced HSP70, likely through binding to HSP70 and affecting its stability, we reasoned that OLA1 and CHIP might compete for binding with HSP70. It is known that CHIP binds to the C-terminus of HSP70.34, 35 To map the OLA1 binding domain on HSP70 protein, plasmid constructs containing the full-length and truncated mutant HSP70 and the YFP-OLA1 vector were co-overexpressed in HEK-293T cells, and the Myc-tagged HSP70 proteins/fragments were immunoprecipitated with anti-Myc antibodies. Similarly to full-length HSP70, the C-terminal fragment of HSP70 was able to pull down YFP-OLA1 (Figures 6a and b). However, the N-terminal fragment of HSP70 that only contains the ATPase domain could not pull down OLA1 and neither could a longer fragment containing both the ATPase domain and the peptide binding domain but with the variable domain truncated (Figure 6b). This suggests that the C-terminal variable domain of HSP70 is the OLA1 binding site. Moreover, endogenous OLA1 was also found to interact with the C-terminal variable domain of HSP70 when Myc-tagged HSP70 proteins/fragments were ectopically expressed in HEK-293T cells (Figures 6a and c).
OLA1 binds to the C-terminal variable domain of HSP70. (a) Schematic diagram of the full-length HSP70 protein (HSP70-FL) and the three truncated segments. The OLA1 binding capability with these HSP70 segments is summarized on the right hand side of the diagram. ‘+’ represents binding, whereas ‘−’ represents lack of binding. (b) Co-transfection assay. HEK-293T cells were co-transfected with the YFP-OLA1 plasmid and one of the Myc-tagged HSP70 constructs (full length or truncated mutants). Forty-eight hours after the transfection, cells lysates were immunoprecipitated with anti-Myc antibody and the immunopellets were detected by immunoblot analysis with anti-YFP and anti-Myc antibodies. (c) Binding with endogenous OLA1. HEK-293T cells were transiently transfected with the full-length Myc-HSP70 or each of the truncated mutants. Two days later, the cell lysates were immunoprecipitated with the anti-Myc antibody, and the precipitated endogenous OLA1 was probed with anti-OLA1 antibody. The same immunopellets were also probed with anti-Myc antibody (upper panel). The whole cell lysates were immunoblotted by anti-OLA1 and anti-Myc antibodies (lower panel)
OLA1 blocks CHIP-binding site on HSP70
Since OLA1 binds to the C-terminal of HSP70, the same binding site as for CHIP, we tested whether OLA1 and CHIP compete for binding to HSP70. Immunoprecipitation and immunoblot analysis using HEK-293T cells transfected with increasing concentrations of YFP-OLA1 construct revealed that the binding between CHIP and HSP70 was markedly decreased with increasing concentrations of the fusion protein (Figure 7a). These results suggest that OLA1 competes with CHIP for HSP70 binding. Conversely, when HSP70 was immunoprecipitated in Ola1−/− MEFs, more CHIP was co-precipitated than in the control MEFs (Figure 7b), indicating that more CHIP binds to HSP70 in the absence of OLA1. It is noteworthy that no differences in total CHIP protein level were observed between the Ola1 knockout and wild-type MEFs (Figure 7b).
OLA1 competes with CHIP in binding with HSP70 and prevents HSP70 from ubiquitination. (a) Binding of CHIP and HSP70 at the presence of OLA1. HEK-293T cells were transiently transfected with the YFP-OLA1 expression vector in a dose-dependent manner. Forty-eight hours after the transfection, cell lysates were immunoprecipitated with anti-HSP70 antibody or IgG, and the resulting immunopellets were immunoblotted with HSP70, CHIP, or YFP antibodies. (b) Binding of CHIP and HSP70 in the absence of Ola1. Cell lysates prepared from Ola−/− and wild-type MEFs were immunoprecipitated with anti-HSP70 antibody and immunoblotted with HSP70 or CHIP antibodies. The presence of HSP70, CHIP, and OLA1 in the whole cell lysates was also verified by immunoblotting (lower panel). β-Actin was used as the loading control. (c–e) Identification of HSP70/OLA1 and HSP70/CHIP complexes. Endogenous OLA1 (c), HSP70 (d), or CHIP (e) in HEK-293T cells was immunoprecipitated with anti-OLA1, anti-HSP70, or anti-CHIP antibody, respectively. The immunoprecipitates were analyzed by immunoblotting with anti-OLA1, anti-HSP70, and anti-CHIP antibodies. The whole cell lysates were also immunoblotted with anti-OLA1, anti-HSP70, and anti-CHIP antibodies. (f) Ubiquitination of HSP70 is attenuated by overexpression of OLA1. HEK-293T cells were co-transfected with the Myc-tagged HSP70, and HA-tagged ubiquitin, and/or YFP-OLA1 constructs. Two days after the transfection, cells were treated with proteasome inhibitor MG132 (10 μM) for 4 h, and the cell lysates were immunoprecipitated with anti-Myc or anti-HA antibody. The resulted immunopellets were then detected by immunoblot analysis with anti-HA or anti-Myc antibody, reciprocally. Whole cell lysates were also immunoblotted by anti-ubiquitin, anti-Myc, anti-YFP, and anti-β-actin antibodies (lower panel). β-Actin served as a loading control. (g) Ubiquitination of HSP70 is enhanced in the absence of OLA1. Cell lysates from the wild-type (WT) and Ola1−/− (KO) MEFs treated with MG132 or the vehicle were immunoprecipitated with anti-HSP70 antibody. The immunopellets were then detected by immunoblot analysis with anti-ubiquitin antibodies. Whole cell lysates were also analyzed for the presence of HSP70, ubiquitin, and OLA1 using the corresponding antibodies. β-Actin was used as the loading control
We next sought to determine whether HSP70 shuttles between two intracellular complexes, a complex with CHIP and another with OLA1. To probe this question, different antibodies (anti-OLA1, anti-HSP70 and anti-CHIP) were used to immunoprecipitate the HSP70 complexes containing either OLA1 or CHIP, and the components of the complexes were analyzed. We found that OLA1 and CHIP were in the anti-HSP70 immunoprecipitants, OLA1 and HSP70 (but not CHIP) were in the anti-OLA1 immunoprecipitants, and CHIP and HSP70 (but not OLA1) were in the anti-CHIP immunoprecipitants (Figures 7c–e), suggesting that the HSP70/OLA1 and HSP70/CHIP complexes coexist in cells.
Because OLA1 blocks CHIP binding to HSP70, we reasoned that OLA1 might prevent CHIP from ubiquitinating HSP70. HEK-293T cells were co-transfected with HA-ubiquitin, Myc-HSP70, and/or YFP-OLA1 plasmids, and after treatment with proteasome inhibitor MG132, the expressed Myc-tagged HSP70 and HA-tagged ubiquitin were immunoprecipitated with anti-Myc and anti-HA antibodies, respectively. The immunoprecipitates were reciprocally immunoblotted with anti-HA and Myc antibodies to monitor the ubiquitination levels of HSP70. As shown in Figure 7f, poly-ubiquitinated HSP70 was markedly decreased in the cells co-transfected with YFP-OLA1 as compared with the cells without YFP-OLA transfection, suggesting that overexpression of OLA1 prevents HSP70 from ubiquitination. Furthermore, when the endogenously expressed HSP70 in Ola1−/− and the wild-type MEFs was examined with immunoprecipitation, poly-ubiquitinated HSP70 was greatly increased in Ola1-knockout MEFs compared with that in the wild-type MEFs in either the absence or the presence of MG132 (Figure 7g). These results further support the hypothesis that OLA1 not only blocks CHIP binding to HSP70 but also dampens its function in degradation of HSP70, leading to accumulation of intracellular HSP70.
Discussion
In this study, we demonstrated that OLA1 is a positive regulator of the heat-shock response. Knockdown of OLA1, or knockout of Ola1, increased heat shock-induced cell death while overexpression of OLA1 rendered cells more resistant to heat shock (Figures 1 and 2; Supplementary Figures S1 and S2). Moreover, we revealed that OLA1 binds to the carboxyl terminus variable domain of HSP70 and subsequently inhibits CHIP-mediated ubiquitination of HSP70, resulting in increased stability of HSP70 (Figures 3, 4, 5, 6, 7; Supplementary Figure S3).
OLA1 belongs to the TRAFAC class, Obg family, and YchF subfamily of P-loop NTPases, and is highly conserved from bacteria to human,23, 26, 32 pointing to its essential functions in fundamental cellular processes. Protein structure analyses suggested that OLA1 functions as a regulatory protein that interacts with downstream effector protein(s) and exerts its downstream functions by switching between the ADP- and ATP-bound forms.27 The human OLA1 protein is a 45-kDa cytosolic protein ubiquitously expressed in most tissues.33 Interestingly, in an earlier study, we reported that OLA1 suppresses the antioxidant response via non-transcriptional mechanisms.33 The fact that OLA1 has important roles in both heat-shock and antioxidant responses indicates that OLA1 is an essential factor for intracellular homeostasis. All currently known heat-shock response modulators are either factors that act on the HSF1/HSE pathway, or that function as molecular chaperones or co-chaperones.5, 19 Our findings suggest that OLA1 may represent a novel type of heat-shock regulator that modulates HSP70’s protein stability. Since we have found that OLA1 co-immunoprecipitates with HSP70 but is absent in the HSP70/CHIP complex (Figures 7c–e), further studies are warranted to determine whether OLA1 itself is a molecular chaperone or a specific co-chaperone for HSP70.
In this study, we also demonstrated a unique mechanism by which OLA1 upregulates HSP70 to maintain its physiological levels. OLA1 deficiency has no effect on the activity of HSF1 (Supplementary Figure S5), nor on the transcription of HSP70 mRNA at either basal conditions or under heat shock (Supplementary Figure S4). On the other hand, the half-life of HSP70 was significantly shortened in cells with knockdown or knockout of OLA1 (Figures 5a and b) and HSP70 protein was more stable in cells with normal OLA1 levels compared with OLA1-deficient cells (Figure 5c). Therefore, we reasoned that OLA1 might instead affect protein stability. CHIP is an E3 ligase that post-translationally regulates HSP70.18, 22, 34, 35 It is proposed that HSP70 acts as an adapter for CHIP to ubiquitinate chaperone clients.18, 36 Because OLA1 blocks CHIP binding to HSP70 (Figures 7a and b), OLA1 may act on CHIP ubiquitinating chaperone clients by disrupting the binding of the adapter (HSP70) and E3 ligase (CHIP). It will be necessary to study the fate of the HSP70 client proteins in the presence and absence of OLA1 in future studies. Nevertheless, our data have shown that OLA1 competes with CHIP for binding to HSP70 (Figures 7a and b) and levels of HSP70 poly-ubiquitination are inversely correlated with the OLA1 expression (Figures 7f and g). Thus, we present a model for how OLA1 and CHIP cooperatively regulate intracellular HSP70 levels (Figure 8). As has been previously described, CHIP binds to HSP70 and brings ubiquitin chains to HSP70, leading to subsequent degradation through the ubiquitin/proteasome pathway. However, based on the present study, a new cellular factor, OLA1, is able to compete with CHIP for binding with the HSP70 C-terminal variable domain, causing CHIP to disassociate from HSP70, and thus attenuating HSP70 ubiquitination and degradation.
A model of how OLA1 stabilizes HSP70 and protects it from proteasome-mediated degradation. As has been illustrated in previous studies, the E3 ligase, CHIP, binds to HSP70 and brings ubiquitin chain to HSP70. Polyubiquitinated HSP70 is then recognized by proteasome for degradation. In the present study, we demonstrate that OLA1 competes with CHIP in binding to the C-terminal variable domain of HSP70. Once OLA1 binds to HSP70, CHIP loses its access to HSP70 and fails to serve as an E3 ligase to mediate the ubiquitination and degradation of HSP70
Our novel findings on increased stability of HSP70 caused by OLA1 may be highly relevant to the molecular pathogenesis of various diseases. HSP70 has been implicated in the molecular pathogenesis of several neurodegenerative diseases, such as Alzheimer’s disease, Parkinson’s disease, and Huntington’s disease,11, 12, 13, 14 a group of diseases characterized by the progressive loss of structure or function of neurons. Along with the degeneration of neurons, one of the morphological features associated with many of these diseases is the presence of insoluble protein aggregates. Due to the fact that high expression of HSP70 protects neurons against cellular toxicity,11, 12, 13, 14 it is plausible that OLA1 might also protect neurons and thus prevent neurodegenerative diseases by enhancing HSP70 levels and function.
Recently, it has been reported that OLA1 was overexpressed in multiple human malignancies, and our own studies also showed that knockdown of OLA1 expression inhibits the motility and invasion of breast cancer cells.31, 32 Given the important roles of OLA1 in stress responses, it is reasonable to suggest that OLA1 may represent a potential therapeutic target for cancer. One of the hallmarks of cancer cells is the acquired or de novo resistance to diverse stresses such as hypoxia and heat shock.37 It is now well established that both HSP70 and HSP90 are upregulated in various human malignancies, and are often associated with poor prognosis and increased resistance to chemotherapies and radiotherapies. More importantly, recent studies have demonstrated that a chemical inhibitor of HSP70 exerts prominent tumor-selective cytotoxic effects.38, 39 Since we have found that knockdown of OLA1 sensitize cancer cells to heat shock in vitro (Figure 2b; Supplementary Figure S1), it is plausible to hypothesize that targeting OLA1 could compromise the thermotolerance of a growing tumor in vivo, sensitizing the tumors to thermal ablation or other therapies.
In summary, we have unraveled a new mechanism by which HSP70 is regulated at the protein turnover level. We found that a cytosolic ATPase, OLA1, protects cells from heat shock-induced cell death, at least in part, by binding to the C-terminal of HSP70 and preventing it from CHIP-mediated ubiquitination and degradation. Further studies on the molecular functions of OLA1 and its interaction with HSP70 should provide additional insights into the pathogenesis of diseases such as protein misfolding disorders and cancer, and warrant the development of potential therapies targeting the OLA1–HSP70 axis.
Materials and Methods
Antibodies and reagents
Mouse monoclonal antibodies against HSP70 (for immunoprecipitation) (ADI-SPA-810), HSP110 (ADI-SPA-1101), HSP90 (ADI-SPA-830), HSP27 (ADI-SPA-800), and HSF1 (ADI-SPA-901) were purchased from Enzo Life Sciences, Farmingdale, NY, USA. Another monoclonal antibody against HSP70 (for immunoblotting) (MAB1663) was purchased from R&D Systems, Minneapolis, MN, USA. Monoclonal antibodies against FLAG tag (129K4754) and β-actin (012M4821) were obtained from Sigma-Aldrich (St Louis, MO, USA). Monoclonal antibodies against c-Myc tag (sc-40), HA tag (sc-7392) and protein A/G (sc-2001, sc-2002) were from Santa Cruz Biotechnology (Dallas, TX, USA). Monoclonal antibody against CHIP (2080S) and polyclonal antibody against ubiquitin (3933BC) were from Cell Signaling Technology, Danvers, MA, USA. Polyclonal antibodies against OLA1 (ab51077) and GFP/YFP (ab6556) were from Abcam, Cambridge, MA, USA. Secondary antibodies, including anti-mouse IgG peroxidase linked whole antibody (NXA931) and anti-rabbit IgG peroxidase linked whole antibody (NA934V) were from GE Healthcare, Pittsburgh, PA, USA. Proeasome inhibitor MG132 (474790-10MG) was obtained from EMD Millipore, Billerica, MA, USA. All other chemicals used in this study were purchased from Sigma-Aldrich, except otherwise stated.
Cell culture
HEK-293, HeLa, and MDA-MB-231 cells were obtained from ATCC and cultured in Dulbecco’s Modified Eagle’s medium (DMEM, Sigma-Aldrich) containing 10% fetal bovine serum (FBS, Thermo Scientific, Hudson, NH, USA), 100 units/ml penicillin and 100 mg/ml streptomycin (Lonza, Alpharetta, GA, USA) at 37°C with 5% CO2. An Ola1-knockout mouse line was custom made, through Texas A&M Institute for Genomic Medicine (TIGM, Houston, Texas), based on high-throughput retroviral gene trapping technology.40 This specific mouse line was generated from embryonic stem (ES) cells carrying a retroviral vector insertion (OmniBank VICTR48, Lexicon Pharmaceuticals, The Woodlands, TX, USA) at the fifth coding exon of the Ola1 gene. The mutant mice were bred in the 129sv/C57BL6 genetic background. Phenotypic characterization of these mutant mice will be published elsewhere. MEFs were derived from 12.5 day embryos according to standard procedures.41 Immortalized Ola1 (+/+) and Ola1 (−/−) MEF lines were established by repetitive subculturing of the primary MEFs for >30 passages.
Expression constructs and short-hairpin RNA constructs
The cDNA fragments encoding full-length human HSP70 and truncated HSP70 proteins including aa1–441, aa1–508, and aa442–641 were constructed into the mammalian expression vector pcDNA3.1 (Invitrogen, Grand Island, NY, USA) with a C-terminal Myc tag. Full-length human OLA1 cDNA (NM_013341.3) was cloned into either the pCMV-Tag1 vector (Stratagene, Santa Clara, CA, USA) with an N-terminal FLAG tag, or the pdEYFP-N1gen plasmid with a C-terminal YFP tag as previously described.33 The pcDNA3.1-HA-CHIP and HA/Flag-CHIP constructs were kindly provided by Dr. Junn Yanagisawa in University of Tsukuba, Ibaraki, Japan.42 The pcDNA3.1-HA-USP4 and HA-ubiquitin constructs were kindly provided by Dr. Jianhua Yang in Baylor College of Medicine, Houston, USA.43 For generation of the HSP70 promoter-dependent luciferase reporter constructs (PGL4-hsp70-luc), a 426-bp human Hsp70 promoter sequence44 was inserted into the HindIII/XhoI sites of the PGL4.18 [lup2P/Neo] vector (Promega, Madison, WI, USA). The CMV promoter-dependent Renilla luciferase reporter plasmid was purchased from Promega. Sh-RNA plasmids used for targeted silencing of OLA1 (sh-OLA1) and the control non-targeting plasmid (sh-control) were made by inserting the following short-hairpin sequences into the pLVTHM vector (Addgene: #12247): 5′-CCGGGAGGAAATGATTGGGCCCATTCTCGAGAATGGGCCCAATCATTTCCTCTTTTTTG-3′ for sh-OLA1 and 5′-CCGGCAACAAGATGAAGAGCACCAACTCGAGTTGGTGCTCTTCATCTTGTTGTTTTTG-3′ for sh-control. All expression vectors were confirmed by sequencing and purified by a plasmid preparation kit (Macherey Nagel, MN, Bethlehem, PA, USA).
For transfection of the DNA vectors, transfection reagents FuGen 6 (Roche, Florence, SC, USA) or Lipofectamine 2000 (Invitrogen) were used according to manufacturer’s instructions.
siRNA-mediated gene knockdown
Human OLA1 cDNA (NM_013341.3)-specific siRNA (SASI_Hs01_00244684) and the control siRNA (MISSION siRNA Universal Negative Control #1 SIC001) were acquired from Sigma-Aldrich. Cells seeded in 6-well plate were transiently transfected with 5 μM siRNA with the DharmaFECT1 siRNA Transfection Reagent (Thermo Scientific) (#T-2001-03) according to manufacturer’s instructions.
Establishment of the stable OLA1 knockdown MDA-MB-231 cell lines
The pLVTHM sh-Control and sh-OLA1 vectors were transfected into MDA-MB-231 cells. Single-cell clones expressing the respective shRNAs linked with the GFP marker were selected by serial dilution, and were subcultured for >1 month. The knockdown efficiency of the target gene was verified by western blot analysis.
Immunoprecipitation, western blot, and mass spectrometry
Cells were washed with ice-cold PBS (pH 7.4) and then lysed in the lysis buffer (Cell Signaling Technology) containing 25 mM Tris-HCl (pH 7.4), 150 mM NaCl, 1 mM EDTA, 1% NP-40, 5% glycerol supplemented with complete protease inhibitor cocktail (Thermo Scientific) on ice for 30 min. The lysates were centrifuged for 15 min at 4°C. Proteins were immunoprecipitated with the indicated antibodies for 3 h. Protein A/G agarose (Santa Cruz Biotechnology) was then incubated with immunocomplexes overnight and washed six times with ice-cold lysis buffer. For western blot analysis, the cell lysate (50 μg) or immunoprecipitates was heated at 100°C for 10 min in sample loading buffer (Thermo Scientific) and separated by SDS-PAGE (Thermo Scientific, Pierce, Hudson, NH, USA). Proteins were transferred onto polyvinylidene difluoride (PVDF, Bio-Rad, Hercules, CA, USA). The membranes were blocked with 5% non-fat milk for 1 h, probed with primary antibodies at 4°C overnight and incubated with secondary antibody (GE Healthcare). Immunoreactive bands were visualized with a chemiluminescence detection system (Thermo Scientific, Pierce). For Mass Spectrometry analysis, HEK-293T cells were transfected with the FLAG-OLA1 vector or the empty vector. The immunoprecipitates were obtained by using anti-Flag antibody from cell lysates. The target binds were excised and the proteins were identified with Mass Spectrometry operated by the Proteomics Core at The Methodist Hospital Research Institute.
Immunofluorescence cell staining
Cells were grown on sterile glass coverslips, fixed in 4% paraformaldehyde for 15 min, permeabilized by 0.2% Triton X-100, and then blocked with 1% BSA for 45 min. Antibodies against OLA1 and HSP70 were incubated overnight at 4°C. After washing three times, the cells were probed with fluorescent conjugated secondary antibodies (Invitrogen) for 1 h at room temperature, followed by blue nuclear counterstaining with Hoechst (Invitrogen). Coverslips were mounted and observed with an Olympus Fluoview FV 1000 Laser Scan Confocal Microscope, Olympus America INC, Center Valley, PA, USA. Images were captured using FC-10-ASW 3.0 software, Olympus America INC.
In vitro binding assay
HSP70 protein (Assay Designs) and the recombinant HIS-tagged human OLA1 protein (custom made by Epoch Life Science, Missouri City, TX, USA) were co-incubated at 4°C for 3 h in 20 μl buffer (25 mM Tris-HCl (pH 7.4), 150 mM NaCl, 1 mM EDTA, 1% NP-40, 5% glycerol). The reacted system was amplified using the same buffer, combined with first antibody, and incubated with protein A/G agarose overnight at 4°C, then washed six times with the same buffer followed by SDS-PAGE analysis to identify the binding partner. The amount of recombinant HSP70 and OLA1 proteins used in this assay were verified by Coomassie blue (Invitrogen) staining.
Quantitative RT-PCR analysis
Total RNA was extracted using the TRIZOL reagent (Invitrogen). RT-PCR was carried out with the iScript one-step RT-PCR kit with SYBR Green reagent (Bio-Rad) in a Miniopticon thermal cycler (Bio-Rad) following manufacturer’s instructions. The mRNA level was analyzed by the relative quantification method based the Ct values obtained from the gene of interest and β-actin (the internal control). The primers were designed using Primer3 software (http://frodo.wi.mit.edu/) as following: human HSP70, sense: 5′-GGCGTGATGACTGCCCTGAT-3′, anti-sense: 5′-CGTCCTCCGCTTTGTACTTCT-3′; mouse Hsp70, sense: 5′-GGGCTTTATCTTCCCTGTTA-3′, anti-sense: 5′-TCACCTCCAAGTTCACCAA-3′; human OLA1: sense: 5′-TGGAGAAGTATGACCCAGGT-3′, anti-sense: 5′-GCTCGAAACCCAGCCTTAATG-3′; mouse Ola1: sense: 5′-TGGAAGATTTGGAACCTCACTGA-3′, anti-sense 5′-AGAATGGGAAGTTTTCTGCTGAA-3′; human β-actin: sense: 5′-CATGTACGTTGCTATCCAGGC-3′, anti-sense: 5′-CTCCTTAATGTCACGCACGAT-3′; mouse β-actin: sense: 5′-TTGCTGACAGGATGCAGAAG-3′, anti-sense: 5′-AAGGGTGTAAAACGCAGCTC-3′.
HSP70 promoter reporter assay
We used a dual luciferase reporter assay system (Promega) for the HSP70 promoter-dependent reporter assay according to manufacturers’ instruction. Briefly, the PGL4-hsp70-luc and the Renilla luciferase control constructs were transiently co-transfected into cells. Forty-eight hours after transfection, cells were exposed to the indicated conditions and lysed for the luciferase assays. Relative HSP70 promoter-dependent activity was normalized to Renilla luciferase activity. A Monolight 3010 luminometer (BD PharMingen, San Jose, CA, USA) was used to measure the values.
Cell viability assay
Growing cells were treated with the indicated conditions. Cell viability was evaluated either by Trypan blue exclusion assay using a cell counter (Nexcelom Cellometer T4, New England BioGroup, Atkinson, NH, USA), or by the MTS-based CellTiter 96 Aqueous One Solution Reagent (Promega).
Statistics
Statistical analysis was performed by a two-tailed Student’s t-test. Values of P<0.05 were considered to be significant.
Abbreviations
- OLA1:
-
Obg-like ATPase 1
- HSP:
-
heat-shock protein
- HSP70:
-
heat-shock protein 70
- CHIP:
-
C-terminus of Hsp70-binding protein
- HSF1:
-
heat-shock transcription factor 1
- HSE:
-
heat-shock element
- UPS:
-
ubiquitin-proteasome system
- MEFs:
-
mouse embryonic fibroblasts
- WT:
-
wild-type
- KO:
-
knockout
- RT-PCR:
-
reverse transcription polymerase chain reaction
- CHX:
-
cycloheximide
- MTS:
-
3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium
References
Lindquist S . The heat-shock response. Annu Rev Biochem 1986; 55: 1151–1191.
Sorger PK . Heat shock factor and the heat shock response. Cell 1991; 65: 363–366.
Mukhopadhyay I, Nazir A, Saxena DK, Chowdhuri DK . Heat shock response: hsp70 in environmental monitoring. J Biochem Mol Toxicol 2003; 17: 249–254.
Richter K, Haslbeck M, Buchner J . The heat shock response: life on the verge of death. Mol Cell 2010; 40: 253–266.
Anckar J, Sistonen L . Regulation of HSF1 function in the heat stress response: implications in aging and disease. Annu Rev Biochem 2011; 80: 1089–1115.
Vabulas RM, Raychaudhuri S, Hayer-Hartl M, Hartl FU . Protein folding in the cytoplasm and the heat shock response. Cold Spring Harb Perspect Biol 2010; 2: a004390.
Mayer MP, Bukau B . Hsp70 chaperones: cellular functions and molecular mechanism. Cell Mol Life Sci 2005; 62: 670–684.
Liberek K, Lewandowska A, Zietkiewicz S . Chaperones in control of protein disaggregation. EMBO J 2008; 27: 328–335.
Daugaard M, Rohde M, Jaattela M . The heat shock protein 70 family: Highly homologous proteins with overlapping and distinct functions. FEBS Lett 2007; 581: 3702–3710.
Rudiger S, Buchberger A, Bukau B . Interaction of Hsp70 chaperones with substrates. Nat Struct Biol 1997; 4: 342–349.
Stetler RA, Gan Y, Zhang W, Liou AK, Gao Y, Cao G et al. Heat shock proteins: cellular and molecular mechanisms in the central nervous system. Prog Neurobiol 2010; 92: 184–211.
Yoo BC, Seidl R, Cairns N, Lubec G . Heat-shock protein 70 levels in brain of patients with Down syndrome and Alzheimer’s disease. J Neural Transm Suppl 1999; 57: 315–322.
Fiszer U, Fredrikson S, Czlonkowska A . Humoral response to hsp 65 and hsp 70 in cerebrospinal fluid in Parkinson’s disease. J Neurol Sci 1996; 139: 66–70.
McLear JA, Lebrecht D, Messer A, Wolfgang WJ . Combinational approach of intrabody with enhanced Hsp70 expression addresses multiple pathologies in a fly model of Huntington’s disease. FASEB J 2008; 22: 2003–2011.
Xanthoudakis S, Nicholson DW . Heat-shock proteins as death determinants. Nat Cell Biol 2000; 2: E163–E165.
Jaattela M . Heat shock proteins as cellular lifeguards. Ann Med 1999; 31: 261–271.
Mosser DD, Morimoto RI . Molecular chaperones and the stress of oncogenesis. Oncogene 2004; 23: 2907–2918.
Qian SB, McDonough H, Boellmann F, Cyr DM, Patterson C . CHIP-mediated stress recovery by sequential ubiquitination of substrates and Hsp70. Nature 2006; 440: 551–555.
Silver JT, Noble EG . Regulation of survival gene hsp70. Cell Stress Chaperones 2012; 17: 1–9.
Chehna-Patel N, Warty N, Sachdeva G, Khole V . Proteolytic tailoring of the heat shock protein 70 and its implications in the pathogenesis of endometriosis. Fertil Steril 2011; 95: 1560–1567 e1–3.
Wang XS, Shankar S, Dhanasekaran SM, Ateeq B, Sasaki AT, Jing X et al. Characterization of KRAS rearrangements in metastatic prostate cancer. Cancer Discov 2011; 1: 35–43.
Kundrat L, Regan L . Identification of residues on Hsp70 and Hsp90 ubiquitinated by the cochaperone CHIP. J Mol Biol 2010; 395: 587–594.
Leipe DD, Wolf YI, Koonin EV, Aravind L . Classification and evolution of P-loop GTPases and related ATPases. J Mol Biol 2002; 317: 41–72.
Paduch M, Jelen F, Otlewski J . Structure of small G proteins and their regulators. Acta Biochim Pol 2001; 48: 829–850.
Sprang SR . G protein mechanisms: insights from structural analysis. Annu Rev Biochem 1997; 66: 639–678.
Czyz A, Wegrzyn G . The Obg subfamily of bacterial GTP-binding proteins: essential proteins of largely unknown functions that are evolutionarily conserved from bacteria to humans. Acta Biochim Pol 2005; 52: 35–43.
Koller-Eichhorn R, Marquardt T, Gail R, Wittinghofer A, Kostrewa D, Kutay U et al. Human OLA1 defines an ATPase subfamily in the Obg family of GTP-binding proteins. J Biol Chem 2007; 282: 19928–19937.
Guerrero C, Milenkovic T, Przulj N, Kaiser P, Huang L . Characterization of the proteasome interaction network using a QTAX-based tag-team strategy and protein interaction network analysis. Proc Natl Acad Sci USA 2008; 105: 13333–13338.
Guerrero C, Tagwerker C, Kaiser P, Huang L . An integrated mass spectrometry-based proteomic approach: quantitative analysis of tandem affinity-purified in vivo cross-linked protein complexes (QTAX) to decipher the 26 S proteasome-interacting network. Mol Cell Proteomics 2006; 5: 366–378.
Gradia DF, Rau K, Umaki AC, de Souza FS, Probst CM, Correa A et al. Characterization of a novel Obg-like ATPase in the protozoan Trypanosoma cruzi. Int J Parasitol 2009; 39: 49–58.
Sun H, Luo X, Montalbano J, Jin W, Shi J, Sheikh MS et al. DOC45, a novel DNA damage-regulated nucleocytoplasmic ATPase that is overexpressed in multiple human malignancies. Mol Cancer Res 2010; 8: 57–66.
Zhang JW, Rubio V, Zheng S, Shi ZZ . Knockdown of OLA1, a regulator of oxidative stress response, inhibits motility and invasion of breast cancer cells. J Zhejiang Univ Sci B 2009; 10: 796–804.
Zhang J, Rubio V, Lieberman MW, Shi ZZ. OLA1, an Obg-like ATPase, suppresses antioxidant response via nontranscriptional mechanisms. Proc Natl Acad Sci USA 2009; 106: 15356–15361.
Jiang J, Ballinger CA, Wu Y, Dai Q, Cyr DM, Hohfeld J et al. CHIP is a U-box-dependent E3 ubiquitin ligase: identification of Hsc70 as a target for ubiquitylation. J Biol Chem 2001; 276: 42938–42944.
Ballinger CA, Connell P, Wu Y, Hu Z, Thompson LJ, Yin LY et al. Identification of CHIP, a novel tetratricopeptide repeat-containing protein that interacts with heat shock proteins and negatively regulates chaperone functions. Mol Cell Biol 1999; 19: 4535–4545.
Wiederkehr T, Bukau B, Buchberger A . Protein turnover: a CHIP programmed for proteolysis. Curr Biol 2002; 12: R26–R28.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011; 144: 646–674.
Galluzzi L, Giordanetto F, Kroemer G . Targeting HSP70 for cancer therapy. Mol Cell 2009; 36: 176–177.
Leu JI, Pimkina J, Frank A, Murphy ME, George DL . A small molecule inhibitor of inducible heat shock protein 70. Mol Cell 2009; 36: 15–27.
Zambrowicz BP, Friedrich GA, Buxton EC, Lilleberg SL, Person C, Sands AT . Disruption and sequence identification of 2000 genes in mouse embryonic stem cells. Nature 1998; 392: 608–611.
Hogan B, Beddington R, Costantini F, Lacy E . Manipulating the Mouse Embryo: A Laboratory Manual 2nd ed. Cold Spring Harbor Laboratory Press: New York, 1994.
Tateishi Y, Kawabe Y, Chiba T, Murata S, Ichikawa K, Murayama A et al. Ligand-dependent switching of ubiquitin-proteasome pathways for estrogen receptor. EMBO J 2004; 23: 4813–4823.
Fan YH, Yu Y, Mao RF, Tan XJ, Xu GF, Zhang H et al. USP4 targets TAK1 to downregulate TNFα-induced NF-κB activation. Cell Death Differ 2011; 18: 1547–1560.
Wu BJ, Kingston RE, Morimoto RI . Human HSP70 promoter contains at least two distinct regulatory domains. Proc Natl Acad Sci USA 1986; 83: 629–633.
Acknowledgements
We are grateful to Dr. Brian E. O’Neill (The Methodist Hospital) for assistance preparing the manuscript. We thank Dr. Junn Yanagisawa (University of Tsukuba) for providing CHIP expression constructs. We thank Dr. Sean Yu (Epoch Life Science) for critical help in cloning of DNA constructs, expression of recombinant OLA1 protein, and purification of anti-human OLA1 antibody and Dr. Jiawei Zhang (Zhejiang University) for establishing the stable knockdown cell lines. We thank Ms. Sunae Kim for assistance in ordering reagents. Intracellular co-localization imaging was performed at The Methodist Hospital Research Institute (TMHRI) Advanced Cellular and Tissue Microscope Core Facility. This work was supported by NIH grant R01CA155069 (ZS) and TMHRI Scholar Award (ZS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by A Stephanou
Supplementary Information accompanies the paper on Cell Death and Disease website
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Mao, RF., Rubio, V., Chen, H. et al. OLA1 protects cells in heat shock by stabilizing HSP70. Cell Death Dis 4, e491 (2013). https://doi.org/10.1038/cddis.2013.23
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2013.23
Keywords
This article is cited by
-
Inhibition of BETs prevents heat shock-induced cell death via upregulating HSPs in SV40 large T antigen transfected cells
Genes & Genomics (2022)
-
Splicing reprogramming of TRAIL/DISC-components sensitizes lung cancer cells to TRAIL-mediated apoptosis
Cell Death & Disease (2021)
-
Heat shock response regulates stimulus-specificity and sensitivity of the pro-inflammatory NF-κB signalling
Cell Communication and Signaling (2020)
-
The molecular chaperone Hsp70 from the thermotolerant Diptera species differs from the Drosophila paralog in its thermostability and higher refolding capacity at extreme temperatures
Cell Stress and Chaperones (2019)
-
Metabolic responses to ethanol and butanol in Chlamydomonas reinhardtii
Biotechnology for Biofuels (2017)