Abstract
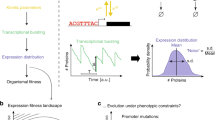
Random fluctuations in genetic networks are inevitable as chemical reactions are probabilistic and many genes, RNAs and proteins are present in low numbers per cell. Such ‘noise’ affects all life processes and has recently been measured using green fluorescent protein (GFP). Two studies show that negative feedback suppresses noise, and three others identify the sources of noise in gene expression. Here I critically analyse these studies and present a simple equation that unifies and extends both the mathematical and biological perspectives.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Schroedinger, E. What is Life? (Cambridge Univ. Press, Cambridge, 1944)
Novick, A. & Weiner, M. Enzyme induction as an all-or-none phenomenon. Proc. Natl Acad. Sci. USA 43, 553–566 (1957)
Maloney, P. C. & Rotman, B. Distribution of suboptimally induced β-d-galactosidase in Escherichia coli. The enzyme content of individual cells. J. Mol. Biol 73, 77–91 (1973)
Løbner-Olesen, A. Distribution of minichromosomes in individual Escherichia coli cells: implications for replication control. EMBO J. 18, 1712–1721 (1999)
Becskei, A. & Serrano, L. Engineering stability in gene networks by autoregulation. Nature 405, 590–593 (2000)
Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002)
Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 183–186 (2002)
Blake, W. J., Kærn, M., Cantor, C. R. & Collins, J. J. Noise in eukaryotic gene expression. Nature 422, 633–637 (2003)
Xie, X. S. Single-molecule approach to dispersed kinetics and dynamic disorder: Probing conformational fluctuation and enzymatic dynamics. J. Chem. Phys. 117, 11024–11032 (2002)
Zwanzig, R. Nonequilibrium Statistical Mechanics (Oxford Univ. Press, New York, 2001)
van Kampen, N. G. Stochastic Processes in Physics and Chemistry (North-Holland, Amsterdam, 1992)
Elf, J., Paulsson, J., Berg, O. G. & Ehrenberg, M. Near-critical phenomena in intracellular metabolite pools. Biophys. J. 84, 154–170 (2003)
Keizer, K. Statistical Thermodynamics of Nonequilibrium Processes (Springer, Berlin, 1987)
Elf, J. & Ehrenberg, M. Fast evaluations of fluctuations in biochemical networks with the linear noise approximation. Genome Res. 13, 2475–2484 (2003)
Kacser, H. & Burns, J. A. The control of flux. Symp. Soc. Exp. Biol. 27, 65–104 (1973)
Westerhoff, H. V. & Van Dam, K. Thermodynamics and Control of Biological Free Energy Transduction (Elsevier, Amsterdam, 1987)
Savageau, M. A. Biochemical Systems Analysis (Addison-Wesley, Reading, 1976)
Savageau, M. A. Parameter sensitivity as a criterion for evaluating and comparing the performance of biochemical systems. Nature 229, 542–544 (1971)
Paulsson, J. & Ehrenberg, M. Noise in a minimal regulatory network: plasmid copy number control. Q. Rev. Biophys. 34, 1–59 (2001)
Dogterom, M. & Leibler, S. Physical aspects of the growth and regulation of microtubule structures. Phys. Rev. Lett. 70, 1347–1350 (1993)
Berg, O. G., Paulsson, J. & Ehrenberg, M. Fluctuations and quality of control in biological cells: zero-order ultrasensitivity reinvestigated. Biophys. J. 79, 1228–1236 (2000)
Rigney, D. R. & Schieve, W. C. Stochastic model of linear, continuous protein-synthesis in bacterial populations. J. Theor. Biol. 69, 761–766 (1977)
Berg, O. G. A model for statistical fluctuations of protein numbers in a microbial-population. J. Theor. Biol. 173, 307–320 (1978)
McAdams, H. H. & Arkin, A. Stochastic mechanisms in gene expression. Proc. Natl Acad. Sci. USA 94, 814–819 (1997)
Paulsson, J., Berg, O. G. & Ehrenberg, M. Stochastic focusing: fluctuation-enhanced sensitivity of intracellular regulation. Proc. Natl Acad. Sci. USA 97, 7148–7153 (2000)
Thattai, M. & van Oudenaarden, A. Intrinsic noise in gene regulatory networks. Proc. Natl Acad. Sci. USA 98, 8614–8619 (2001)
Peccoud, J. & Ycart, B. Markovian modelling of gene-product synthesis. Theor. Popul. Biol. 48, 222–234 (1995)
Kepler, T. B. & Elston, T. C. Stochasticity in transcriptional regulation: origins, consequences, and mathematical representations. Biophys. J. 81, 3116–3136 (2001)
Sasai, M. & Wolynes, P. G. Stochastic gene expression as a many-body problem. Proc. Natl Acad. Sci. USA 100, 2374–2379 (2003)
Singh, U. N. Polyribosomes and unstable messenger RNA: a stochastic model of protein synthesis. J. Theor. Biol. 25, 444–460 (1969)
Swain, P. S., Elowitz, M. B. & Siggia, E. D. Intrinsic and extrinsic contributions to stochasticity in gene expression. Proc. Natl Acad. Sci. USA 99, 12795–12800 (2002)
Savageau, M. A. Comparison of classical and autogenous systems of regulation in inducible operons. Nature 252, 546–549 (1974)
Thieffry, D., Huerta, A. M., Perez-Rueda, E. & Collado-Vides, J. From specific gene regulation to genomic networks: a global analysis of transcriptional regulation in Escherichia coli. Bioessays 20, 433–440 (1998)
Shen-Orr, S. S., Milo, R., Mangan, S. & Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nature Genet. 31, 64–68 (2002)
Paulsson, J. Multileveled selection on plasmid replication. Genetics 161, 1373–1384 (2002)
Chattoraj, D. K. Control of plasmid DNA replication by iterons: no longer paradoxical. Mol. Microbiol. 37, 467–476 (2000)
Vilar, J. M., Kueh, H. Y., Barkai, N. & Leibler, S. Mechanisms of noise-resistance in genetic oscillators. Proc. Natl Acad. Sci. USA 99, 5988–5992 (2002)
Howard, M. & Rutenberg, A. D. Pattern formation inside bacteria: fluctuations due to the low copy number of proteins. Phys. Rev. Lett. 90, 128102 (2003)
Acknowledgements
This work was supported by a Lewis-Thomas fellowship from Princeton University and by the Cambridge-MIT Institute. I thank A. Becskei, E. Cox, M. Ehrenberg, J. Elf, M. Elowitz, I. Golding, S. Leibler, A. Løbner-Olesen, C. Peterson, S. Sawai, P. Swain, M. Thattai and A. van Oudenaarden for discussions and comments on the manuscript. I also thank W. Blake, M. Kaern, C. Cantor and J. Collins for sharing unpublished data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares that he has no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Paulsson, J. Summing up the noise in gene networks. Nature 427, 415–418 (2004). https://doi.org/10.1038/nature02257
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02257
This article is cited by
-
Design principles of improving the dose-response alignment in coupled GTPase switches
npj Systems Biology and Applications (2023)
-
Flagellar dynamics reveal fluctuations and kinetic limit in the Escherichia coli chemotaxis network
Scientific Reports (2023)
-
Transcriptional noise adjusted for expression levels reveals genes with high transcriptional noise that are highly expressed, functionally related, and co-regulated in yeast
Current Genetics (2022)
-
Propagation of stochastic fluctuations in RNA polymerase abundance in constitutive gene expression
Journal of the Korean Physical Society (2022)
-
Single-cell measurement of plasmid copy number and promoter activity
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.