Abstract
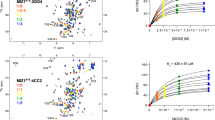
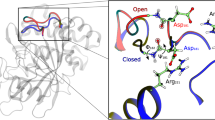
Signal transduction, regulatory processes and pharmaceutical responses are highly dependent upon ligand residence times. Gaining insight into how physical factors influence residence times (1/koff) should enhance our ability to manipulate biological interactions. We report experiments that yield structural insight into koff involving a series of eight 2,4-diaminopyrimidine inhibitors of dihydrofolate reductase whose binding affinities vary by six orders of magnitude. NMR relaxation-dispersion experiments revealed a common set of residues near the binding site that undergo a concerted millisecond-timescale switching event to a previously unidentified conformation. The rate of switching from ground to excited conformations correlates exponentially with the binding affinity Ki and koff, suggesting that protein dynamics serves as a mechanical initiator of ligand dissociation within this series and potentially for other macromolecule-ligand systems. Although the forward rate of conformational exchange, kconf,forward, is faster than koff, the use of the ligand series allowed for connections to be drawn between kinetic events on different timescales.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Tummino, P.J. & Copeland, R.A. Residence time of receptor-ligand complexes and its effect on biological function. Biochemistry 47, 5481–5492 (2008).
Lu, H. & Tonge, P.J. Drug-target residence time: critical information for lead optimization. Curr. Opin. Chem. Biol. 14, 467–474 (2010).
Olson, J.S. & Phillips, G.N. Jr. Kinetic pathways and barriers for ligand binding to myoglobin. J. Biol. Chem. 271, 17593–17596 (1996).
Bourgeois, D. et al. Complex landscape of protein structural dynamics unveiled by nanosecond Laue crystallography. Proc. Natl. Acad. Sci. USA 100, 8704–8709 (2003).
Bourne, H.R., Sanders, D.A. & McCormick, F. The GTPase superfamily: conserved structure and molecular mechanism. Nature 349, 117–127 (1991).
Ishima, R., Freedberg, D.I., Wang, Y.X., Louis, J.M. & Torchia, D.A. Flap opening and dimer-interface flexibility in the free and inhibitor-bound HIV protease, and their implications for function. Structure 7, 1047–1055 (1999).
Eisenmesser, E.Z. et al. Intrinsic dynamics of an enzyme underlies catalysis. Nature 438, 117–121 (2005).
Boehr, D.D., McElheny, D., Dyson, H.J. & Wright, P.E. The dynamic energy landscape of dihydrofolate reductase catalysis. Science 313, 1638–1642 (2006).
Labeikovsky, W., Eisenmesser, E.Z., Bosco, D.A. & Kern, D. Structure and dynamics of pin1 during catalysis by NMR. J. Mol. Biol. 367, 1370–1381 (2007).
Masterson, L.R. et al. Dynamics connect substrate recognition to catalysis in protein kinase A. Nat. Chem. Biol. 6, 821–828 (2010).
Sapienza, P.J., Mauldin, R.V. & Lee, A.L. Multi-timescale dynamics study of FKBP12 along the rapamycin-mTOR binding coordinate. J. Mol. Biol. 405, 378–394 (2010).
Mauldin, R.V., Carroll, M.J. & Lee, A.L. Dynamic dysfunction in dihydrofolate reductase results from antifolate drug binding: modulations of dynamics within a structural state. Structure 17, 386–394 (2009).
Carroll, M.J. et al. Direct detection of structurally resolved dynamics in a multi-conformation receptor-ligand complex. J. Am. Chem. Soc. 133, 6422–6428 (2011).
Eisenmesser, E.Z., Bosco, D.A., Akke, M. & Kern, D. Enzyme dynamics during catalysis. Science 295, 1520–1523 (2002).
Zolli-Juran, M., Cechetto, J.D., Hartlen, R., Daigle, D.M. & Brown, E.D. High throughput screening identifies novel inhibitors of Escherichia coli dihydrofolate reductase that are competitive with dihydrofolate. Bioorg. Med. Chem. Lett. 13, 2493–2496 (2003).
Matthews, D.A. et al. Dihydrofolate reductase: X-ray structure of the binary complex with methotrexate. Science 197, 452–455 (1977).
Sawaya, M.R. & Kraut, J. Loop and subdomain movements in the mechanism of Escherichia coli dihydrofolate reductase: crystallographic evidence. Biochemistry 36, 586–603 (1997).
Loria, J.P., Rance, M. & Palmer, A.G. A relaxation-compensated Carr-Purcell-Meiboom-Gill sequence for characterizing chemical exchange by NMR spectroscopy. J. Am. Chem. Soc. 121, 2331–2332 (1999).
Boehr, D.D., McElheny, D., Dyson, H.J. & Wright, P.E. Millisecond timescale fluctuations in dihydrofolate reductase are exquisitely sensitive to the bound ligands. Proc. Natl. Acad. Sci. USA 107, 1373–1378 (2010).
Palmer, A.G. III, Kroenke, C.D. & Loria, J.P. Nuclear magnetic resonance methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol. 339, 204–238 (2001).
Osborne, M.J., Venkitakrishnan, R.P., Dyson, H.J. & Wright, P.E. Diagnostic chemical shift markers for loop conformation and substrate and cofactor binding in dihydrofolate reductase complexes. Protein Sci. 12, 2230–2238 (2003).
Mulder, F.A., Mittermaier, A., Hon, B., Dahlquist, F.W. & Kay, L.E. Studying excited states of proteins by NMR spectroscopy. Nat. Struct. Biol. 8, 932–935 (2001).
Baccanari, D.P. & Joyner, S.S. Dihydrofolate reductase hysteresis and its effect of inhibitor binding analyses. Biochemistry 20, 1710–1716 (1981).
Appleman, J.R., Howell, E.E., Kraut, J., Kuhl, M. & Blakley, R.L. Role of aspartate 27 in the binding of methotrexate to dihydrofolate reductase from Escherichia coli. J. Biol. Chem. 263, 9187–9198 (1988).
Namanja, A.T. et al. Toward flexibility-activity relationships by NMR spectroscopy: dynamics of Pin1 ligands. J. Am. Chem. Soc. 132, 5607–5609 (2010).
Frederick, K.K., Marlow, M.S., Valentine, K.G. & Wand, A.J. Conformational entropy in molecular recognition by proteins. Nature 448, 325–329 (2007).
Berger, A. & Linderstrom-Lang, K. Deuterium exchange of poly-dl-alanine in aqueous solution. Arch. Biochem. Biophys. 69, 106–118 (1957).
Hvidt, A. & Nielsen, S.O. Hydrogen exchange in proteins. Adv. Protein Chem. 21, 287–386 (1966).
Hansen, D.F., Vallurupalli, P. & Kay, L.E. Using relaxation dispersion NMR spectroscopy to determine structures of excited, invisible protein states. J. Biomol. NMR 41, 113–120 (2008).
Hansen, D.F., Vallurupalli, P., Lundstrom, P., Neudecker, P. & Kay, L.E. Probing chemical shifts of invisible states of proteins with relaxation dispersion NMR spectroscopy: how well can we do? J. Am. Chem. Soc. 130, 2667–2675 (2008).
Gangjee, A., Zaveri, N., Queener, S.F. & Kisliuk, R.L. Synthesis and biological activities of tetrahydroquinazoline analogs of aminopterin and methotrexate. J. Heterocycl. Chem. 32, 243–247 (1995).
Wider, G. & Dreier, L. Measuring protein concentrations by NMR spectroscopy. J. Am. Chem. Soc. 128, 2571–2576 (2006).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on Unix pipes. J. Biomol. NMR 6, 277–293 (1995).
Johnson, B.A. & Blevins, R.A. NMRView: a computer program for the visualization and analysis of NMR data. J. Biomol. NMR 4, 603–614 (1994).
Skrynnikov, N.R., Dahlquist, F.W. & Kay, L.E. Reconstructing NMR spectra of “invisible” excited protein states using HSQC and HMQC experiments. J. Am. Chem. Soc. 124, 12352–12360 (2002).
Summerfield, R.L. et al. A 2.13 A structure of E. coli dihydrofolate reductase bound to a novel competitive inhibitor reveals a new binding surface involving the M20 loop region. J. Med. Chem. 49, 6977–6986 (2006).
Fierke, C.A., Johnson, K.A. & Benkovic, S.J. Construction and evaluation of the kinetic scheme associated with dihydrofolate reductase from Escherichia coli. Biochemistry 26, 4085–4092 (1987).
Acknowledgements
The authors thank G. Young (University of North Carolina at Chapel Hill Biomolecular NMR Facility) and K. Koshlap (University of North Carolina at Chapel Hill Eshelman School of Pharmacy NMR Facility) for their excellent technical assistance. We also acknowledge H. Kohn for assistance in assigning and interpreting the small-molecule NMR spectra as well as N. Kett for chiral resolution of 3 and the lab of P. Guengerich for use of their stopped-flow fluorimeter. M.J.C. gratefully acknowledges predoctoral fellowships from the ACS Division of Medicinal Chemistry (supported by Pfizer Global R&D), the American Foundation for Pharmaceutical Education (supported by the Rho Chi Society and Schering-Plough) and the Graduate School at UNC. This work was funded by US National Institutes of Health grant GM083059 to A.L.L.
Author information
Authors and Affiliations
Contributions
M.J.C., A.L.L. and S.F.S. designed research; M.J.C., R.V.M., A.V.G. and E.J.C. performed research; M.J.C., R.V.M., A.V.G., S.F.S., E.J.C. and A.L.L. analyzed data; and M.J.C. and A.L.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 13890 kb)
Rights and permissions
About this article
Cite this article
Carroll, M., Mauldin, R., Gromova, A. et al. Evidence for dynamics in proteins as a mechanism for ligand dissociation. Nat Chem Biol 8, 246–252 (2012). https://doi.org/10.1038/nchembio.769
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.769
This article is cited by
-
The role of NMR in leveraging dynamics and entropy in drug design
Journal of Biomolecular NMR (2020)
-
Dynamic regulation of GDP binding to G proteins revealed by magnetic field-dependent NMR relaxation analyses
Nature Communications (2017)
-
The drug–target residence time model: a 10-year retrospective
Nature Reviews Drug Discovery (2016)
-
Allosteric Effect of Adenosine Triphosphate on Peptide Recognition by 3′5′-Cyclic Adenosine Monophosphate Dependent Protein Kinase Catalytic Subunits
The Protein Journal (2016)
-
A conserved P-loop anchor limits the structural dynamics that mediate nucleotide dissociation in EF-Tu
Scientific Reports (2015)