Abstract
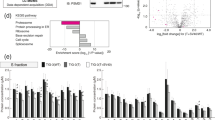
The ubiquitin/proteasome pathway has been implicated in a wide variety of cellular processes and the number of substrates degraded by the proteasome is impressive. Most prominently, the stability of a large number of transcription factors is regulated by ubiquitination. To elucidate pathways regulated by the proteasome, gene expression profiles were generated, comparing changes of mRNA expression of 7900 genes from the UniGene collection upon exposure of cells to the proteasome inhibitors Lactacystin, Lactacystin-β-lactone or MG132 by means of microarray based cDNA hybridization. The three profiles were very similar, but differed significantly from a gene expression profile generated with the histone deacetylase inhibitor Trapoxin A, indicating that the observed alterations were indeed due to proteasome inhibition. Two of the most prominently induced genes encoded the growth arrest and DNA damage inducible protein Gadd153 and the activating transcription factor ATF3, both transcription factors of the CCAAT/enhancer binding protein (C/EBP) family. A third gene encoded for the transcriptional repressor and c-Myc antagonist Mad1. Our results suggest that proteasome inhibition leads to upregulation of specific members of transcription factor families controlling cellular stress response and proliferation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Adams J, Palombella VJ, Sausville EA, Johnson J, Destree A, Lazarus DD, Maas J, Pien CS, Prakash S and Elliott PJ . 1999 Cancer Res 59: 2615–2622
Amundson SA, Zhan Q, Penn LZ and Fornace Jr AJ . 1998 Oncogene 17: 2149–2154
Amundson SA, Bittner M, Chen Y, Trent J, Meltzer P and Fornace Jr AJ . 1999 Oncogene 18: 3666–3672
Ayer DE, Kretzner L and Eisenman RN . 1993 Cell 72: 211–222
Bai JP . 1995 J Pharm Pharmacol 47: 674–677
Blagosklonny MV, Wu GS, Omura A and el Deiry WS . 1996 Biochem Biophys Res Commun 227: 564–569
Bogyo M, Gaczynska M and Ploegh HL . 1997 Biopolymers 43: 269–280
Bonvini P, Nguyen P, Trepel J and Neckers LM . 1998 Oncogene 16: 1131–1139
Bush KT, Goldberg AL and Nigam SK . 1997 J Biol Chem 272: 9086–9092
Cailleau R, Young R, Olive M and Reeves Jr WJ . 1974 J Natl Cancer Inst 53: 661–674
Chang DD, Park N-H, Denny CT, Nelson SF and Pe M . 1998 Oncogene 16: 1921–1930
Chen BP, Wolfgang CD and Hai T . 1996a Mol Cell Biol 16: 1157–1168
Chen C, Nussenzweig A, Guo M, Kim D, Li GC and Ling CC . 1996b Oncogene 17: 1659–1665
Drexler HCA . 1998 Apoptosis 3: 1–7
Eymin B, Dubrez L, Allouche M and Solary E . 1997 Cancer Res 57: 686–695
Fawcett TW, Eastman HB, Martindale JL and Holbrook NJ . 1996 J Biol Chem 271: 14285–14289
Fawcett TW, Martindale JL, Guyton KZ, Hai T and Holbrook NJ . 1999 Biochem J 339: 135–141
Firestein R and Feuerstein N . 1998 J Biol Chem 273: 5892–5902
Giard DJ, Aaronson SA, Todaro GJ, Arnstein P, Kersey JH, Dosik H and Parks WP . 1973 J Natl Cancer Inst 51: 1417–1423
Goodbourn S and King P . 1997 Biochem Soc Transac 25: 498–502
Han S, Park K, Kim HY, Lee MS, Kim HJ and Kim YD . 1999 Oncol Rep 6: 569–574
Heid CA, Stevens J, Livak KJ and Williams PM . 1996 Genomic Res 6: 986–994
Herrmann JL, Briones Jr F, Brisbay S, Logothetis CJ and McDonnell TJ . 1998 Oncogene 17: 2889–2899
Hofmann F, Martelli F, Livingston DM and Wang Z . 1996 Genes Dev 10: 2949–2959
Imajoh-Ohmi S, Kawaguchi T, Sugiyama S, Tanaka K, Omura S and Kikuchi H . 1995 Biochem Biophys Res Commun 217: 1070–1077
Ichihara A and Tanaka K . 1995 Mol Biol Rep 21: 49–52
Kawazoe Y, Nakai A, Tanabe M and Nagata K . 1998 Eur J Biochem 255: 356–362
Kitagawa H, Tani E, Ikemoto H, Ozaki I, Nakano A and Omura S . 1999 FEBS Lett 443: 181–186
Lee DH and Goldberg AL . 1998 Trends Cell Biol 8: 397–403
Liang G, Wolfgang CD, Chen BP, Chen TH and Hai T . 1996 J Biol Chem 271: 1695–1701
Lloyd RV, Erickson LA, Jin L, Kulig E, Qian X, Cheville JC and Scheithauer BW . 1999 Am J Pathol 154: 313–323
Machiels BM, Henfling ME, Gerards WL, Broers JL, Bloemendal H, Ramaekers FC and Schutte B . 1997 Cytometry 28: 243–252
Masumoto M, Minami M, Takeda K, Sakao Y and Akira S . 1996 FEBS Lett 395: 143–147
Miller G, Fuchs R and Lai E . 1997 Genome Res 7: 1027–1032
Outinen PA, Sood SK, Liaw PC, Sarge KD, Maeda N, Hirsh J, Ribeau J, Podor TJ, Weitz JI and Austin RC . 1998 Biochem J 332: 213–221
Palombella VJ, Rando OJ, Goldberg AL and Maniatis T . 1994 Cell 78: 773–785
Paul SR, Bennett F, Calvetti JA, Kelleher K, Wood CR, O'Hara Jr RM, Leary AC, Sibley B, Clark SC, Williams DA and Yang YC . 1990 Proc Natl Acad Sci USA 87: 7512–7516
Roussel MF, Ashmun RA, Sherr CJ, Eisenman RN and Ayer DE . 1996 Mol Cell Biol 16: 2796–2801
Schauber C, Chen L, Tongaonkar P, Vega I, Lambertson D, Potts W and Madura K . 1998 Nature 391: 715–718
Schena M, Shalon D, Davis RW and Brown PO . 1995 Science 270: 467–470
Schena M, Shalon D, Heller R, Chai A, Brown PO and Davis RW . 1996 Proc Natl Acad Sci USA 93: 10614–10619
Shalon D, Smith SJ and Brown PO . 1996 Genome Res 6: 639–645
Talley AK, Dewhurst S, Perry SW, Dollard SC, Gummuluru S, Fine SM, New D, Epstein LG, Gendelman HE and Gelbard HA . 1995 Mol Cell Biol 15: 2359–2366
Tchernitsa OI, Zuber J, Sers C, Brinckmann R, Britsch SK, Adams V and Schäfer R . 1999 Oncogene 18: 5448–5454
User bulletin #2, ABI PRISM 7700 Sequence Detection System 1997 PE Applied Biosystems
Van Lint C, Emiliani S and Verdin E . 1996 Gene Expr 5: 245–253
Wang X, Luo H, Chen H, Duguid W and Wu J . 1998a J Immunol 160: 788–801
Wang XZ, Kuroda M, Sok J, Batchvarova N, Kimmel R, Chung P, Zinszner H and Ron D . 1998b EMBO J 17: 3619–3630
Wang XZ, Lawson B, Brewer JW, Zinszner H, Sanjay A, Mi LJ, Boorstein R, Kreibich G, Hendershot LM and Ron D . 1996 Mol Cell Biol 16: 4273–4280
Wolfgang CD, Chen BP, Martindale JL, Holbrook NJ and Hai T . 1997 Mol Cell Biol 17: 6700–6707
Yoshida H and Sugita K . 1992 Jpn J Cancer Res 83: 324–328
Zinszner H, Kuroda M, Wang X, Batchvarova N, Lightfoot RT, Remotti H, Stevens JL and Ron D . 1998 Genes Dev 12: 982–985
Acknowledgements
We are grateful to the staff at Incyte Inc. (Palo Alto, CA, USA) for performing the array based hybridizations. We thank Lidia Sambucetti for Trapoxin A and Johannes Roesel for his constructive comments on the manuscript.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Zimmermann, J., Erdmann, D., Lalande, I. et al. Proteasome inhibitor induced gene expression profiles reveal overexpression of transcriptional regulators ATF3, GADD153 and MAD1. Oncogene 19, 2913–2920 (2000). https://doi.org/10.1038/sj.onc.1203606
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1203606
Keywords
This article is cited by
-
ATF3 induction prevents precocious activation of skeletal muscle stem cell by regulating H2B expression
Nature Communications (2023)
-
Phase 1 study of ixazomib, an investigational proteasome inhibitor, in advanced non-hematologic malignancies
Investigational New Drugs (2015)
-
Gambogic acid is cytotoxic to cancer cells through inhibition of the ubiquitin-proteasome system
Investigational New Drugs (2013)
-
Indole-3-carbinol synergistically sensitises ovarian cancer cells to bortezomib treatment
British Journal of Cancer (2012)
-
Microarray profiling reveals the integrated stress response is activated by halofuginone in mammary epithelial cells
BMC Research Notes (2011)