Abstract
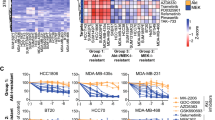
The multidrug resistance (MDR) phenotype is a major cause of cancer treatment failure. Here the expressions of 4224 genes were analysed for association with intrinsic or acquired doxorubicin (DOX) resistance. A cluster of overexpressed genes related to DOX resistance was observed. Included in this cluster was ABCB1 the P-glycoprotein transporter protein gene and MMP1 (Matrix Metalloproteinase 1), indicative of the invasive nature of resistant cells, and the oxytocin receptor (OXTR), a potential new therapeutic target. Overexpression of genes associated with xenobiotic transformation, cell transformation, cell signalling and lymphocyte activation was also associated with DOX resistance as was estrogen receptor negativity. In all carcinoma cells, compared with HBL100 a putatively normal breast epithelial cell line, a cluster of overexpressed genes was identified which included several keratins, in particular keratins 8 and 18 which are regulated through the ras signalling pathway. Analysis of genomic amplifications and deletions revealed specific genetic alterations common to both intrinsic and acquired DOX resistance including ABCB1, PGY3 (ABCB4) and BAK. The findings shown here indicate new possibilities for the diagnosis of DOX resistance using gene expression, and potential novel therapeutic targets for pharmacological intervention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, Boldrick JG, Sabet H, Tran T, Yu X, Powell JI, Yang LM, Marti GE, Moore T, Hudson J, Lu LS, Lewis DB, Tibshirani R, Sherlock G, Chan WC, Greiner TC, Weisenburger DD, Armitage JO, Warnke R, Levy R, Wilson W, Grever MR, Byrd JC, Botstein D, Brown PO, Staudt LM . 2000 Nature 403: 503–511
Bamberger AN, Methner C, Lisbola BW, Stadtler C, Schulte HM, Loning T, Milde-Langosch K . 1999 Int. J. Cancer 84: 533
Batist G, Tulpule A, Sinha BK, Katki AG, Myers C, Cowan KH . 1986 J. Biol. Chem. 261: 15544–15549
Benbow U, Schoenermark MP, Mitchell TI, Rutter JL, Shimokawa K, Nagase H, Brinckerhoff CE . 1999 J. Biol. Chem. 274: 25371–25378
Blatchford DR, Quarrie LH, Tonner E, McCarthy C, Flint DJ, Wilde CJ . 1999 J. Cell. Physiol. 181: 304–311
Brockstedt E, Rickers A, Kostka S, Laubershiemer A, Dorken B, WittmannLiebold B, Bommert K, Otto A . 1998 J. Biol. Chem. 273: 28057–28064
Cassoni P, Sapina A, Fortunati N, Munaron L, Chini B, Bussolati G . 1997 Int. J. Cancer 72: 340–344
Cassoni P, Fulcheri E, Carcangiu ML, Stella A, Deaglio S, Bussolati G . 2000 J. Path. 190: 470–477
Chang K, Pastan I . 1996 Proc. Natl. Acad. Sci. USA 93: 136–140
Chen G, Jaffrezou JP, Fleming WH, Duran GE, Sikic BI . 1994 Cancer Res. 54: 4980–4987
Davies R, Budworth J, Riley J, Snowden R, Gescher A, Gant TW . 1996 Br. J. Cancer 73: 307–315
DeRisi JL, Iyer VR, Brown PO . 1997 Science 278: 680–686
Eisen MB, Spellman PT, Brown PO, Botstein D . 1998 Proc. Natl. Acad. Sci. USA 95: 14863–14868
Eisen MB, Brown PO . 1999 DNA Arrays for the Analysis of Gene Expression: cDNA Preparation and Characterization. Weissman SM (ed) Academic press: San Diego pp. 179–205
Fairchild CR, Kao-Shan C-S, Wang-Peng J, Rosen N, Israel MA, Malera PW, Cowan KH, Goldsmith ME . 1987 Cancer Res. 47: 5141–5148
Feinberg AP, Vogelstein B . 1983 Anal. Biochem. 132: 6–13
Gottesman MM, Pastan I . 1993 Ann. Rev. Biochem. 62: 385–427
Hanahan D, Weinberg RA . 2000 Cell 100: 57–70
Jones JL, Walker RA . 1997 Int. J. Oncol. 11: 609–616
Klucken J, Buchler C, Orso E, Kaminski WE, PorschOzcurumez M, Liebisch C, Kapinsky M, Diederich W, Drobnik W, Dean M, Allikmets R, Schmitz G . 2000 Proc. Natl. Acad. Sci. USA 97: 817–822
Lala P, Ito S, Lingwood CA . 2000 J. Biol. Chem. 275: 6246–6251
Martin K, Kritzman DM, Price LM, Koh B, Kwan CP, Zhang XH, Mackay A, OHare MJ, Kaelin DM, Mutter GL, Pardee AB, Sager R . 2000 Cancer Res. 60: 2232–2238
Oshima RG, Baribault H, Caulin C . 1996 Cancer Meta. Rev. 15: 445
Perou CM, Jeffrey SS, VandeRijn M, Rees CA, Eisen MB, Ross DT, Pergamenschikov A, Williams CF, Zhu SX, Lee JCF, Lashkari D, Shalon D, Brown PO, Botstein D . 1999 Proc. Natl. Acad. Sci. USA 96: 9212–9217
Perou C, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Perganmenschikov A, Williams C, Zhu S, Lonning PE, Borresen-Dale A-L, Brown PO, Botstein D . 2000 Nature 406: 747–752
Robinson LJ, Roberts WK, Tao Tao Ling. Lamming D, Stemberg SS, Roepe PD . 1997 Biochemistry 36: 11169–11178
Shen D-W, Fojo A, Chin JE, Roninson IB, Richert N, Pastan I, Gottesman MM . 1986 Science 232: 643–645
Stephen R, Corcoran D, Darbre PD . 1998 J. Molec. Endrocrinol. 20: 375–380
Van der Bliek AM, Baas F, Van der Velde-Koerts T, Biedler JL, Meyers MB, Ozolls RF, Hamilton TC, Joenje H, Borst P . 1988 Cancer Res. 48: 5927–5932
Vasudevan S, Tsuruo T, Rose DR . 1998 J. Biol. Chem. 273: 25413–25419
Weinstein RS, Jakate SM, Dominguez JM, Lebovitz MD, Koukoulis GK, Kuszak JR, Klusens LF, Grogan TM, Saclarides TJ, Roninson IB, Coon JS . 1991 Cancer Res. 51: 2720–2726
Acknowledgements
The authors would like to thank Drs M Eisen and J DeRisi for their generous provision of software, Profs G Cohen, R Walker and Drs B Coyle and P Glynn for critical reading of the manuscript Dr K Lilley and colleagues for sequencing and David Jones and colleagues in the University MSB workshops for arrayer construction. TW Gant thanks Drs CA Shari, E Nuwaysir and Ms K Hayes for initial training at NIEHS. Particular thanks are due to Prof G Cohen (deputy director of the MRC Toxicology Unit) for his support of the microarray project.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Turton, N., Judah, D., Riley, J. et al. Gene expression and amplification in breast carcinoma cells with intrinsic and acquired doxorubicin resistance. Oncogene 20, 1300–1306 (2001). https://doi.org/10.1038/sj.onc.1204235
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1204235
Keywords
This article is cited by
-
The oxytocin receptor signalling system and breast cancer: a critical review
Oncogene (2020)
-
Prospect and challenge of detecting dynamic gene copy number increases in stem cells by whole genome sequencing
Journal of Molecular Medicine (2019)
-
SPIN1, negatively regulated by miR-148/152, enhances Adriamycin resistance via upregulating drug metabolizing enzymes and transporter in breast cancer
Journal of Experimental & Clinical Cancer Research (2018)
-
Genetic background influences susceptibility to chemotherapy-induced hematotoxicity
The Pharmacogenomics Journal (2018)
-
Amplification of LAPTM4B and YWHAZ contributes to chemotherapy resistance and recurrence of breast cancer
Nature Medicine (2010)