Abstract
Aims/hypothesis
Metformin is widely used as a hypoglycaemic reagent for type 2 diabetes. While the reduction of hepatic gluconeogenesis is thought to be a key effect, the detailed molecular mechanism of action of metformin remains to be elucidated. To gain insight into this, we performed a global gene expression profiling study.
Materials and methods
We performed DNA microarray analysis to study global gene expression in the livers of obese diabetic db/db mice 2 h after a single administration of metformin (400 mg/kg).
Results
This analysis identified 14 genes that showed at least a 1.5-fold difference in expression following metformin treatment, including a reduction of glucose-6-phosphatase gene expression. The mRNA levels of glucose-6-phosphatase showed one of the best correlations with blood glucose levels among 12,000 genes. Enzymatic activity of glucose-6-phosphatase was also reduced in metformin-treated liver. Moreover, intensive analysis of the expression profile revealed that metformin effected significant alterations in gene expression across at least ten metabolic pathways, including those involved in glycolysis-gluconeogenesis, fatty acid metabolism and amino acid metabolism.
Conclusions/interpretation
These results suggest that reduction of glucose-6-phosphatase activity, as well as suppression of mRNA expression levels of this gene, in liver is of prime importance for controlling blood glucose levels in vivo, at least at early time points after metformin treatment. Our results also suggest that metformin not only affects expression of specific genes, but also alters the expression level of multiple genes linked to the metabolic pathways involved in glucose and lipid metabolism in the liver.
Similar content being viewed by others
Introduction
Metformin is a widely used hypoglycaemic agent for treatment of type 2 diabetic patients. The action of metformin in ameliorating hyperglycaemia is linked to its ability to reduce hepatic gluconeogenesis and to improve peripheral insulin sensitivity [1, 2]. Metformin promotes glucose uptake in muscle [3, 4] and lowers hepatic glucose production [5, 6]. In addition, metformin improves lipid metabolism, lowering plasma triglycerides [7, 8] and non-esterified fatty acids [9].
A detailed mechanistic basis for the hypoglycaemic action of metformin in diabetic patients remains unclear, although a number of reports concerning the cellular response to and molecular function of metformin have been published. Based on data from a variety of experimental conditions, metformin has been shown to activate AMP-activated protein kinase [10] and inhibit mitochondrial respiratory complex I [11, 12], mitochondrial permeability transition [13] and tyrosine phosphatase activity [14]. Additionally, metformin has been reported to influence hepatic gene expression in cultured hepatocytes, with noted alterations in the expression of particular genes, such as glucose-6-phosphatase (G6pc), glucokinase, and 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 2 (Hmgcs2). These genes, which are related to glycolysis-gluconeogenesis or cholesterol metabolism, were assumed to be critical for the action of metformin [15]. However, detailed studies of the action of metformin in the liver, its major target tissue, remain to be performed. Moreover, the effect of metformin on global gene expression in diabetic livers has still to be elucidated.
DNA microarray technology has been widely used to gain insight into the overall effect on gene expression of various drug treatments and pathologies. Results from such analyses have brought useful information in terms of drug action mechanisms, as well as on the development of diseases [16, 17]. The methods used to extract biological meaning from the extensive gene expression data obtained from DNA microarray analysis study are still in progress; however, approaches using multivariate analysis, such as hierarchical clustering and principal component analysis, have been shown to be useful especially for data showing large differences in expression of numerous genes between samples [18–22]. When the number of differentially expressed genes identified is too numerous to extract an overall interpretation from the biological function of individual genes, genes belonging to each metabolic pathway or a ‘gene ontology’ class are treated as a cohort, and can be used to understand the biological significance of the observed changes of expression [23–26].
Therefore, we performed a DNA microarray analysis experiment in order to study the global effect of metformin on gene expression in the livers of db/db mice [27]. We identified G6PC as a key enzyme in the liver in relation to the mechanism of action of metformin. Our pathway analysis also uncovered alterations of gene expression in certain metabolic pathways related to the action of metformin.
Materials and methods
Animals and drug treatment
We used 8-week-old C57BL/Ksj-db/db Jcl male mice (Clea Japan, Tokyo, Japan) for metformin treatment, as well as for other experimental protocols. Metformin was given orally. For DNA microarray analysis, livers were isolated 2 h after metformin administration. Livers from metformin-treated mice were used for the preparation of total RNA for both DNA microarray and RT-PCR analyses. Livers from non-treated or vehicle-treated mice were processed in a similar manner. Before isolation of livers, blood was collected from each mouse for glucose measurement. Animals were maintained on normal chow (CE-2, 352 kcal/100 g; Clea Japan) on a 12-h light/dark cycle. Metformin (1,1-dimethylbiguanide hydrochloride) was purchased from Sigma–Aldrich (St. Louis, MO, USA). All animal experiments were conducted according to the guidelines of Sumitomo Pharmaceuticals Committee on Animal Research.
Measurement of blood glucose levels
Blood samples were collected from tail veins and blood glucose measured (Glucose CII-Test; Wako Chemical, Osaka, Japan) according to the manufacturer’s protocol.
Isolation of total RNA
Total RNA was extracted from each mouse liver with TRIZOL reagent (Invitrogen, Carlsbad, CA, USA) and purified using RNeasy kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. Total RNA prepared from individual animals was used for DNA microarray analysis, either independently, or, when necessary pooled (mixture of equal amount of total RNA from individual animals in each treatment group).
DNA microarray analysis
DNA microarray analysis using Affymetrix GeneChip technology was performed as described previously [28]. Briefly, 10 μg of total RNA was used as a template for cDNA synthesis. The generation of hybridisation cocktails, hybridisation to DNA microarrays (Murine Genome U74A v2), and fluorescent labelling of complementary RNA were carried out as described in the standard Affymetrix protocol (Affymetrix, Santa Clara, CA, USA) [29]. Fluorescently labelled microarrays were scanned with a laser scanner (Hewlett Packard, Palo Alto, CA, USA). Global normalisation was performed to make the average intensity of all probe sets equal to 100. Detection call, signal intensity, fold change value, change call and detection p value were obtained with default settings. Comparative analysis was performed by Microarray Suite 5.0 (Affymetrix).
Real-time quantitative RT-PCR
PCR was performed with a sequence detector system (ABI PRISM 7700; Applied Biosystems, Foster City, CA, USA). Primers (forward: 5′-AGCCTCCGGAAGTATTGTCTCA-3′ and reverse: 5′-TCCACCCCTAGCCCTTTTAGTAG-3′ for mouse, and forward: 5′-GGCCCTCAACTCCAGCATGT-3′ and reverse: 5′-AGAGATGCAGCAGGCCCAAG-3′ for rat) for G6pc genes were designed by Primer Express (Applied Biosystems). Reverse transcription was performed from 2 μg total RNA from db/db mouse livers with oligo-dT primer, and the first strand cDNAs were synthesised with SuperScript II reverse transcriptase. The quantitative PCR reaction was performed using TaqMan PCR Reagent Kits according to the manufacturer’s protocol (Applied Biosystems). Sample cDNAs equivalent to 100 ng of RNA were examined in each reaction in a 96-well PCR plate. The following temperature profile was used: 50 min at 42°C for reverse transcription and 30 s at 95°C for denaturation of cDNA/RNA hybrid, followed by 40 cycles of 15 s at 95°C and 40 cycles of 60 s at 60°C. Levels of glyceraldehyde-3-phosphate dehydrogenase were measured for each sample, and used as internal standards.
Measurement of glucose-6-phosphatase activity
Activity of G6PC was examined by Glucose C-II Test (Wako) according to the manufacturer’s protocol. Briefly, frozen mouse livers were homogenised in nine volumes of cold medium containing 0.5 mol/l phosphate buffer, pH 7.4, 5 mmol/l EDTA, 5 mmol/l EGTA, 10% glycerol. The homogenates were then centrifuged at 30,000 g for 30 min, the supernatants were diluted two-fold with glycerol and kept at −30°C until assayed. For this 20 μl of diluted sample containing 100 μg protein, 20 μl of 0.3 mol/l glucose-6-phosphate, 10 μl of distilled water and 200 μl of Wako kit reagent were mixed in a 96-well plate. A reagent blank was set with phosphate buffer instead of samples. Increases in absorbance at 500 nm were measured using a spectrophotometer at 30°C. Activity was determined as the change in absorbance from 15 min to 20 min per μg protein.
In vitro assay of metformin-regulated gene expression
Rat H4IIE cells were cultured in alpha-minimal essential medium supplemented with 10% fetal bovine serum, 100 U/ml penicillin, 100 μg/ml streptomycin and 0.25 μg/ml amphotericin B at 37°C in an atmosphere of 5% CO2. Metformin was added to the cells at the indicated concentrations. After metformin treatment cells were used for RT-PCR analysis and the medium was used for measurement of glucose concentration as described above.
Other methods
Detailed bioinformatical and statistical methods for data analysis are available online as Electronic Supplementary Material (ESM).
Results
Blood glucose levels were reduced in diabetic mice by a single administration of metformin
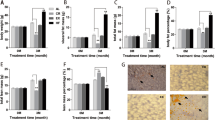
First, we examined blood glucose levels after a single administration of metformin in db/db mice, and observed a reduction of blood glucose levels as shown previously [30]. Metformin (200 and 400 mg/kg) significantly lowered blood glucose levels in mice, even following a single administration of the agent (Fig. 1a). The decrease occurred in a dose-dependent manner 3–6 h after metformin treatment; levels returned to original levels 24 h after drug administration. Having confirmed that a single administration of metformin caused a hypoglycaemic activity in db/db mice, we conducted an analysis designed to elucidate the mechanism of action of metformin in vivo.
Blood glucose levels in db/db mice after metformin administration. a Blood glucose levels in db/db mice treated with 0, 200 and 400 mg/kg metformin. Blood glucose levels were measured at 1, 3, 6, 24 h after administration. Open circles, vehicle alone; diamonds, 200 mg/kg metformin; squares, 400 mg/kg metformin. b Blood glucose levels of mice used in DNA microarray analysis study. Two hours after administration of metformin, the blood glucose levels in the 50 and 400 mg/kg metformin-treated group were 20.0±1.5 and 16.8±1.4 mmol/l, respectively, as compared with 23.1±3.2 and 22.3±2.2 mmol/l for groups without treatment and those treated with vehicle alone. Results are the mean±SD of 12 animals (a) of five animals (b) for each treatment condition. **p<0.01, *p<0.05 when compared with vehicle alone
Change of gene expression in diabetic livers after metformin treatment, as observed by DNA microarray analysis
To gain insight into the molecular mechanism of metformin’s action, we performed a global gene expression profiling study using DNA microarray technology. In more detail, mice were treated with either high-dose (400 mg/kg) or low-dose (50 mg/kg) metformin, or vehicle alone (0.5% methylcellulose). For control purposes, we also examined a group of mice receiving no treatment. Each group contained five animals. Given that significant reductions in blood glucose levels were observed between 1 and 3 h after metformin treatment (Fig. 1a), the RNA used for gene expression profile analysis was derived from the livers of animals killed 2 h after drug treatment. At this particular time point, only high-dose metformin caused a significant reduction in blood glucose levels compared with vehicle alone (Fig. 1b). In contrast, little, if any, reduction was seen in mice treated with low-dose metformin.
Following this confirmation, we performed DNA microarray analysis via Affymetrix GeneChip technology (Murine Genome U74Av2) with total RNA prepared from the livers of these mice. We compared gene expression levels between vehicle and metformin-treated (400 mg/kg) livers using pooled samples, in which equal amounts of RNA from five individual animals were mixed for each group. Surprisingly, among the 12,488 genes represented on the array, only 8 and 25 showed significant changes (increase or decrease call as given by Affymetrix microarray suite software) in expression of more than 2- and 1.5-fold, respectively. We also performed DNA microarray analysis for individual samples, and identified those exhibiting statistically significant differences in expression levels between vehicle and high-dose metformin treatment. In the individual sample analysis, 36 genes showed significant changes (p<0.05), 14 of which showed a more than 1.5-fold difference in expression terms (Table 1). Interestingly, G6pc, whose signal intensity was much higher than any of the other genes listed in Table 1, is the rate-limiting enzyme for gluconeogenesis in the liver. Suppression of G6pc expression by metformin was confirmed by quantitative RT-PCR (Fig. 2a). Furthermore, enzymatic activity of G6PC was also significantly decreased by metformin treatment (Fig. 3). Interestingly, nuclear receptor subfamily 0, group B, member 2 (Nr0b2), previously known as small heterodimer partner (Shp), which was reported to be an important factor linked to expression regulation of gluconeogenesis genes [31], was also included in the list.
Correlation between G6pc expression in the liver and blood glucose levels. a G6pc expression in the livers of db/db mice with no treatment, and 2 h after metformin (50 or 400 mg/kg) or vehicle administration. G6pc expression level was determined by quantitative RT-PCR. Results are the mean±SD of five animals for each treatment condition. **p<0.01 when compared to vehicle alone. b, c Scatter plots indicating relationship between blood glucose concentration and G6pc expression levels for each animal. G6pc expression level was determined by DNA microarray (b) or by quantitative RT-PCR (c) analysis. Pearson’s correlation coefficient values were 0.833 and 0.881 for DNA microarray and quantitative RT-PCR analysis, respectively. Open squares, control; open circles, vehicle alone; closed circles, 50 mg/kg metformin; closed squares, 400 mg/kg metformin
Enzymatic activity of G6PC in the liver after metformin treatment. Enzymatic activity for G6PC was measured in the liver of db/db mice 2 h after vehicle or 400 mg/kg metformin administration. Results are presented as the mean±SD of five animals for each treatment condition. *p<0.05 when compared with vehicle alone
G6pc gene expression was correlated with blood glucose levels and dose of metformin
As demonstrated in Fig. 1b, blood glucose levels showed a dose-dependent reduction in response to metformin. Blood glucose reduction by metformin is attributed to gluconeogenesis in the liver. As mentioned above, one of the major changes following metformin treatment in liver was down-regulation of G6PC, which is the rate limiting enzyme in gluconeogenesis. Accordingly, we assessed correlation of G6pc expression with blood glucose in individual animals by both RT-PCR and microarray analyses. As shown in Fig. 2b,c, there was a clear positive correlation (r>0.8) between these factors. Regulation of G6pc expression in the liver may be of prime importance for controlling blood glucose levels in vivo, at least at early time points after metformin treatment.
Among the 12,488 genes we analysed, a clear positive correlation (r>0.8) with blood glucose level was found for expression levels of only six genes, including G6pc. We examined the reproducibility of our results by an independent experiment, confirming that, among this subset of six genes, only G6pc and the gene encoding solute carrier family 25 (mitochondrial carrier, phosphate carrier), member 25 (Slc25a25) showed a high correlation with blood glucose levels (data not shown). Both of these genes are included in Table 1.
Metformin causes down-regulation of G6pc in vitro
The in vivo experiments described above suggest that down-regulation of G6pc is of prime importance in the context of blood glucose reduction in this model. In order to exclude the possibility that the reduction of G6pc expression was caused by decreased blood glucose levels, we employed a hepatocyte-derived cell line H4IIE. Metformin (1 or 3 mmol/l) was added to the culture medium, and expression of G6pc and medium glucose level were compared with control cells. The results clearly demonstrate that more than 20% down-regulation of G6pc mRNA expression was caused after metformin treatment, and the change is independent of medium glucose level (Fig. 4). These results support our hypothesis that liver G6pc is down-regulated by metformin, and it is a key mechanism in the reduction of blood glucose level following a single administration of the drug.
G6pc expression and glucose concentration in H4IIE cells after metformin treatment. Relative G6pc gene expression levels (closed bars) in H4IIE cells 2 h after metformin (1 or 3 mmol/l) or vehicle treatment were determined by quantitative RT-PCR. Relative glucose concentration (open bars) in the medium of the H4IIE cells was determined as described in Materials and methods. Results are presented as the mean±SD of three times for each treatment. *p<0.05 when compared with vehicle alone
Significant changes in gene expression by metformin treatment were also detected within metabolic pathways related to glucose and lipid metabolism
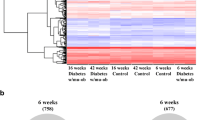
Both the number of differentially expressed genes and the magnitude of the differences observed were smaller than our prior expectations. Expression differences between the two conditions, namely vehicle and 400 mg/kg treatment, were detected for only 72 genes in the pooled sample analysis, even when the magnitude of the difference was not considered. In order to examine the relevance of these subtle gene expression changes, we carried out gene expression profile analysis using materials from 20 individual animals (no treatment, vehicle, 200 and 400 mg/kg metformin; n=5) and performed unsupervised hierarchical clustering of all 20 sets of expression data with the 72 genes. The results clearly showed that animals that had been treated with high-dose metformin and showed significant pharmacological activity were clustered together and could be separated from most of the other animals (Fig. 5a). Furthermore, principal component analysis using the same 72 gene dataset showed a remarkable shift in the distribution of animals treated with high-dose metformin compared with the others (Fig. 5b). This analysis suggests that treatment with 50 mg/kg metformin causes a small degree of gene expression changes, even though the dose was not enough to show a significant pharmacological effect (Fig. 1b). In contrast, the animals receiving either vehicle alone or no treatment were indistinguishable in both analysis methods. It should be noted that: (1) neither animals treated with a low dose of metformin, nor animals without treatment were used to deduce the 72 genes; and (2) most of the 72 genes showed a lower than 1.5-fold difference. These data suggest that the expression difference we detected in this experiment is intrinsic to each treatment, and that the overall tendency of the changes can be described by using many genes simultaneously, even though each expression change looks marginal.
Multivariate analysis of gene expression data. Expression profile data derived from pooled RNA of the livers of animals treated with vehicle alone and of animals treated with 400 mg/kg metformin were compared to each other, with only 72 genes showing significant differences in gene expression levels. Individual samples were analysed by unsupervised hierarchical clustering analysis (a) and principal component analysis (b) using data on the 72 differentially expressed genes. Animals treated with 400 mg/kg metformin were clearly separated from other animals in both analyses. Colour key: a red, increased relative expression; shades of blue, decreased relative expression; b red circles, 400 mg/kg metformin; pink circles, 50 mg/kg metformin; light blue circles, vehicle alone; deep blue circles, no treatment
In order to understand the biological relevance of such slight changes, we considered groups of genes in each metabolic pathway. Substantially affected pathways, if any, should show larger global expression changes than non-relevant pathways. Among 105 known metabolic pathways registered in the Kyoto Encyclopaedia of Genes and Genomes (KEGG) database, 59 pathways composed of more than four genes were chosen for the following analysis. First, the signal intensity for genes in each pathway was subjected to principal component analysis, in which the position of the points in space reflected integrated gene expression profiles derived from each animal. Then, in order to select the pathways changed by administration of metformin, we examined whether the observed distribution was significantly different between animals treated with a high dose of metformin (n=5) and the remaining animals (n=15). Twenty metabolic pathways were chosen by the analysis, which showed differential expression with statistical significance (p<0.01; Wilks’ lambda). A pattern recognition algorithm, namely Support Vector Machine (Visual Mining Studio version 1.4; Mathematical Systems, Tokyo, Japan), was then applied to examine the extent of differential expression that corresponded to the high-dose metformin-treated cohort compared with the other experimental groups. As a result of the analysis, the ten most substantially affected pathways were identified and their names, along with KEGG pathway IDs, are shown in the Textbox.

Although most of the genes in these pathways were not included in the initial list of 72 differentially expressed genes identified from comparison analysis, the derived pathways are closely related to the physiological action of metformin [1, 5, 8]. The pathways mentioned above were not obtained when metformin-treated animals (n=10), including both 50 and 400 mg/kg treatment conditions, were compared with the remaining animal groups, i.e. vehicle alone and no treatment (n=10) (data not shown). This suggests that the observed expression changes are associated with the pharmacological activity of metformin, but are not dependent on administration of the drug. Thus, metformin should affect expression of metabolic pathways involved in glucose and lipid metabolism in diabetic liver even a short time after metformin treatment.
Discussion
Metformin is a widely used hypoglycaemic agent for diabetes, and the reduction of hepatic gluconeogenesis is believed to be a major function of metformin. This is the first report of global gene expression analysis in livers of diabetic mice treated with metformin. We observed down-regulation of G6pc levels, and expression changes of genes in metabolic pathways related to the hypoglycaemic effect of metformin, i.e. not only with respect to individual genes, in the livers of these mice 2 h after 400 mg/kg metformin administration.
As mentioned, we used db/db mice in this study. First, we confirmed the blood glucose lowering effect of metformin in db/db mice. A significant reduction of blood glucose levels was observed in animals treated with a high dose of metformin (400 mg/kg), whereas the group treated with a low dose of metformin (50 mg/kg) tended to have reduced glucose levels that were not statistically significant. This result correlated well with previous studies, which demonstrated that the efficacy dose of metformin in rodent was around 300 mg/kg [32].
Following DNA microarray analysis of individual RNA from livers of groups treated with a high dose of metformin and groups treated with vehicle alone, we detected 14 genes that exhibited a significant change in expression. Among these, G6pc showed the highest signal intensity, suggesting a biological significance of this gene. G6pc has been linked to the action of metformin according to in vitro studies [15, 33] (Fig. 4). Indeed, G6pc showed one of the best correlations with blood glucose levels (Fig. 2b,c) among more than 12,000 genes analysed in this study. Enzymatic activity of G6PC was also reduced by metformin treatment (Fig. 3). Interestingly, even when metformin was administered every day for 4 weeks to db/db mice, a similar correlation between G6pc expression and blood glucose levels was observed in the liver 2 h after the last administration (data not shown). It could be objected that the down-regulation of G6pc mRNA and reduction of blood glucose level occurred too quickly, since the drug was orally administered. However, it has been reported that the blood concentration of metformin reached almost maximal level within 1 h following oral administration in the rat [34], and intraperitoneal administration of an inhibitor of G6PC translocase reduced blood glucose levels significantly 30 min after administration [35]. Furthermore, it has recently been reported that mice injected with short hairpin RNA exhibited a significant reduction in postprandial glucose levels accompanied by a decrease in G6pc expression [36]. Thus, it would be reasonable to suppose that regulation of G6pc gene expression in the liver is of prime importance for controlling blood glucose levels in vivo, at least at early time points, after metformin treatment.
As shown in Fig. 4, in vitro experiment suggests that the down-regulation of G6pc induced by metformin should be independent of the extracellular glucose level. But we could not exclude the possibility that the down regulation is mediated by an intermediate metabolite of gluconeogenesis, since our pathway analysis indicated that a cohort of genes in the glycolysis and gluconeogenesis pathways showed expression changes after metformin treatment. However, since expression of other genes on the pathways did not change to the same large degree as G6pc and only G6pc was highly correlated with blood glucose, the regulation mechanism would be different between G6pc and other genes in the glycolysis and gluconeogenesis pathways.
The mechanisms by which metformin regulates G6pc remain to be elucidated; however, a recent study demonstrated that metformin treatment for 3 h suppressed G6pc expression independently of insulin signalling in rat hepatoma H4IIE cells [33]. G6pc gene expression was observed in an in vivo study to be modulated by polyunsaturated fatty acids [37]. Interestingly, as shown in Table 1, a transcription regulatory protein, Nr0b2, that regulates gluconeogenic genes [31], was down-regulated by metformin treatment, as observed in the case of G6pc. However, the correlation between Nr0b2 expression and blood glucose levels was not significant. It was reported that NR0B2 represses G6pc transcription mediated by the transcription factor forkhead box O1 (FOXO1) by competing with a coactivator [31]. Under our assay conditions, no significant change of expression was observed, either for Foxo1, or for phosphoenolpyruvate carboxykinase, which is also regulated by NR0B2 [31]. Taken together, the down-regulation of G6pc should be mediated by a mechanism independent of NR0B2 and the FOXO1 system. Expression of Slc25a25 also changed considerably and showed a high correlation with blood glucose levels. It is a putative mitochondrial calcium-dependent solute carrier protein, whose function(s) remain to be determined. It might play a role in energy metabolism and/or in the action of metformin, since the gene is highly expressed in liver and skeletal muscle [38]. Interestingly, all the changes listed in Table 1 are down-regulation, which may suggest that these genes were regulated by similar mechanisms.
Our analysis also suggests that the expression of many genes was affected by metformin treatment. By focusing on groups of genes in metabolic pathways, instead of on individual genes, we found it highly likely that changes in mRNA expression occurred in cohorts of genes in certain metabolic pathways, accompanied by reduction of glucose levels by metformin treatment. We also observed that similar serial administration of metformin caused changes in similar metabolic pathways including glycolysis-gluconeogenesis and fatty acid metabolism (data not shown). Changes in the fatty acid and glycerolipid metabolism pathways may contribute to an improvement of lipid metabolism by repeated treatment with metformin, while some fatty acids might be important in modulating mRNA expression of genes such as G6pc [37]. Interestingly, changes of expression in the bile acid biosynthesis, but not the sterol biosynthesis pathway, were detected; indeed, co-ordinated transcriptional regulation between bile acids and gluconeogenesis [39, 40], as well as suppression of gluconeogenenic genes by bile acid [31], have been reported recently. With regard to amino acid metabolism pathways, such as valine, leucine and isoleucine degradation, lysine degradation, arginine and proline metabolism and tryptophan metabolism, it has previously been reported that protein metabolism is abnormal in type 2 diabetes [41], although its relevance to the action of metformin remains to be elucidated.
As shown in Fig. 5, all of the isolated pathways are closely connected to each other around the glucose-energy metabolism pathways, and acetyl coenzyme A and pyruvate should be at the centre of the changes we observed (Fig. 6). The co-ordinate regulation in gene expression for these metabolic pathways in the liver may facilitate improvement of hyperglycaemia via treatment with metformin. Though the regulation mechanism remains to be elucidated, it is possibly of interest that two signalling molecules and two transcription factors are included in Table 1.
Overview of the identified pathways, showing the inter-relationship of the identified metabolic pathways. All pathways are positioned around glucose metabolism pathways, namely glycolysis-gluconeogenesis, the citrate cycle and pyruvate metabolism, and connected to each other via acetyl coenzyme A (Acetyl-CoA) and/or pyruvate
Our global gene expression analysis approach identified significant changes of some metabolic pathways at an early time point, even when changes of individual genes were not significantly detectable. Pharmacological effect through changes of metabolic pathways, and not only with respect to individual genes, could be important for the action of metformin, and this kind of effect on global gene expression might be used in determining the mechanism of action of other drugs.
Since the method we developed, which is called ‘pathway analysis’, does not require prior knowledge on the mechanism of the compound, this approach could be useful with regard to other drugs whose molecular mechanisms of action are unknown, as well as in the search for novel pharmacological activities of compounds of interest.
Abbreviations
- Foxo1:
-
forkhead box O1
- G6pc:
-
glucose-6-phosphatase
- Hmgcs2:
-
3-hydroxy-3-methylglutaryl-Coenzyme A synthase 2
- KEGG:
-
Kyoto Encyclopaedia of Genes and Genomes
- Nr0b2:
-
nuclear receptor subfamily 0, group B, member 2
- Slc25a25:
-
solute carrier family 25 (mitochondrial carrier, phosphate carrier), member 25
References
Bailey CJ, Turner RC (1996) Metformin. N Engl J Med 334:574–579
Kirpichnikov D, McFarlane SI, Sowers JR (2002) Metformin: an update. Ann Intern Med 137:25–33
Hundal HS, Ramlal T, Reyes R, Leiter LA, Klip A (1992) Cellular mechanism of metformin action involves glucose transporter translocation from an intracellular pool to the plasma membrane in L6 muscle cells. Endocrinology 131:1165–1173
Musi N, Hirshman MF, Nygren J et al (2002) Metformin increases AMP-activated protein kinase activity in skeletal muscle of subjects with type 2 diabetes. Diabetes 51:2074–2081
Hundal RS, Krssak M, Dufour S et al (2000) Mechanism by which metformin reduces glucose production in type 2 diabetes. Diabetes 49:2063–2069
Stumvoll M, Nurjhan N, Perriello G, Dailey G, Gerich JE (1995) Metabolic effects of metformin in non-insulin-dependent diabetes mellitus. N Engl J Med 333:550–554
DeFronzo RA, Goodman AM (1995) Efficacy of metformin in patients with non-insulin-dependent diabetes mellitus. N Engl J Med 333:541–549
Cusi K, Consoli A, DeFronzo RA (1996) Metabolic effect of metformin on glucose and lactate metabolism in non-insulin-dependent diabetes mellitus. J Clin Endocrinol Metab 81:4059–4067
Abbasi F, Kamath V, Rizvi AA, Carantoni M, Chen YD, Reaven GM (1997) Results of a placebo-controlled study of the metabolic effects of the addition of metformin to sulfonylurea-treated patients: evidence for a central role of a adipose tissue. Diabetes Care 20:1863–1869
Zhou G, Myers R, Li Y et al (2001) Role of AMP-activated protein kinase in mechanism of metformin action. J Clin Invest 108:1167–1174
Owen MR, Doran E, Halestrap AP (2000) Evidence that metformin exerts its anti-diabetic effects through inhibition of complex 1 of the mitochondrial respiratory chain. Biochem J 348:607–614
El-Mir MY, Nogueira V, Fontaine E, Averet N, Rigoulet M, Leverve X (2000) Dimethylbiguanide inhibits cell respiration via an indirect effect targeted on the respiratory chain complex I. J Biol Chem 275:223–228
Guigas B, Detaille D, Chauvin C et al (2004) Metformin inhibits mitochondrial permeability transition and cell death: a pharmacological in vitro study. Biochem J 82:877–884
Holland W, Morrison T, Chang Y, Wiernsperger N, Stith BJ (2004) Metformin (Glucophage) inhibits tyrosine phosphatase activity to stimulate the insulin receptor tyrosine kinase. Biochem Pharmacol 67:2081–2091
Fulgencio JP, Kohl C, Girard J, Pegorier JP (2001) Effect of metformin on fatty acid and glucose metabolism in freshly isolated hepatocytes and on specific gene expression in cultured hepatocytes. Biochem Pharmacol 62:439–446
Golub TR, Slonim DK, Tamayo P et al (1999) Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286:531–537
Ramaswamy S, Golub TR (2002) DNA microarrays in clinical oncology. J Clin Oncol 20:1932–1941
Butte A (2002) The use and analysis of microarray data. Nat Rev Drug Discov 1:951–960
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Spellman PT, Sherlock G, Zhang MQ et al (1998) Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol Biol Cell 9:3273–3297
Raychaudhuri S, Stuart JM, Altman RB (2000) Principal component analysis to summarize microarray experiments: application to sporulation time series. Pac Symp Biocomput 455–466
Hilsenbeck SG, Friedrichs WE, Schiff R et al (1999) Statistical analysis of array expression data as applied to the problem of tamoxifen resistance. J Natl Cancer Inst 91:453–459
Doniger SW, Salomonis N, Dahlquist KD, Vranizan K, Lawlor SC, Conklin BR (2003) MAPPFinder: using gene ontology and GenMAPP to create a global gene-expression profile from microarray data. Genome Biol 4:R7
Draghici S, Khatri P, Martins RP, Ostermeier GC, Krawetz SA (2003) Global functional profiling of gene expression. Genomics 8:98–104
Grosu P, Townsend JP, Hartl DL, Cavalieri D (2002) Pathway processor: a tool for integrating whole-genome expression results into metabolic networks. Genome Res 12:1121–1126
Mootha VK, Lindgren CM, Eriksson KF et al (2003) PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34:267–273
Chen H, Charlat O, Tartaglia LA et al (1996) Evidence that the diabetes gene encodes the leptin receptor: identification of a mutation in the leptin receptor gene in db/db mice. Cell 84:491–495
Ishida N, Hayashi K, Hoshijima M et al (2002) Large scale gene expression analysis of osteoclastogenesis in vitro and elucidation of NFAT2 as a key regulator. J Biol Chem 277:41147–41156
Lockhart DJ, Dong H, Byrne MC et al (1996) Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat Biotechnol 14:1675–1680
Fujita H, Fujishima H, Morri T et al (2002) Effect of metformin on adipose tissue resistin expression in db/db mice. Biochem Biophys Res Commun 298:345–349
Yamagata K, Daitoku H, Shimamoto Y et al (2004) Bile acids regulate gluconeogenic gene expression via small heterodimer partner-mediated repression of hepatocyte nuclear factor 4 and Foxo 1. J Biol Chem 279:23158–23165
Yasuda N, Inoue T, Nagakura T et al (2002) Enhanced secretion of glucagon-like peptide 1 by biguanide compounds. Biochem Biophys Res Commun 298:779–784
Morioka K, Nakatani K, Matsumoto K et al (2005) Metformin-induced suppression of glucose-6-phosphatase expression is independent of insulin signaling in rat hepatoma cells. Int J Mol Med 15:449–452
Stepensky D, Friedman M, Srour W, Raz I, Hoffman A (2001) Preclinical evaluation of pharmacokinetic-pharmacodynamic rationale for oral CR metformin formulation. J Control Release 71:107–115
Parker JC, VanVolkenburg MA, Levy CB et al (1998) Plasma glucose levels are reduced in rats and mice treated with an inhibitor of glucose-6-phosphatase translocase. Diabetes 47:1630–1636
Huang A, Chen Y, Wang X et al (2004) Functional silencing of hepatic microsomal glucose-6-phosphatase gene expression in vivo by adenovirus-mediated delivery of short hairpin RNA. FEBS Lett 558:69–73
Rajas F, Gautier A, Bady I, Montano S, Mithieux G (2002) Polyunsaturated fatty acyl coenzyme A suppress the glucose-6-phosphatase promoter activity by modulating the DNA binding of hepatocyte nuclear factor 4 alpha. J Biol Chem 277:15736–15744
Mashima H, Ueda N, Ohno H et al (2003) A novel mitochondrial Ca2+-dependent solute carrier in the liver identified by mRNA differential display. J Biol Chem 278:9520–9527
Shin DJ, Campos JA, Gil G, Osborne TF (2003) PGC-1alpha activates CYP7A1 and bile acid biosynthesis. J Biol Chem 278:50047–50052
De Fabiani E, Mitro N, Gilardi F, Caruso D, Galli G, Crestani M (2003) Coordinated control of cholesterol catabolism to bile acids and of gluconeogenesis via a novel mechanism of transcription regulation linked to the fasted-to-fed cycle. J Biol Chem 278:39124–39132
Gougeon R, Pencharz PB, Marliss EB (1994) Effect of NIDDM on the kinetics of whole-body protein metabolism. Diabetes 43:318–328
Acknowledgements
This study was supported by Dainippon Sumitomo Pharma, Japan. We are grateful to A. Nishihara for assistance with animal experiments and to Y. Ujimasa for DNA microarray experiments. We also thank K. Kikuchi, M. Ruto and H. Hanafusa for their valuable advice and assistance.
Duality of interest
All authors are employees of Dainippon Sumitomo Pharma, who manufacture metformin in Japan.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the supplementary material.
Rights and permissions
About this article
Cite this article
Heishi, M., Ichihara, J., Teramoto, R. et al. Global gene expression analysis in liver of obese diabetic db/db mice treated with metformin. Diabetologia 49, 1647–1655 (2006). https://doi.org/10.1007/s00125-006-0271-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-006-0271-y